K-Nearest neighbor is a great technique for dealing with data. In the video below, we will look at how to use this tool with Python.
Category Archives: Statistics
Z-Scores VIDEO
Calculating Z-Scores

Data Exploration Case Study: Credit Default
Exploratory data analysis is the main task of a Data Scientist with as much as 60% of their time being devoted to this task. As such, the majority of their time is spent on something that is rather boring compared to building models.
This post will provide a simple example of how to analyze a dataset from the website called Kaggle. This dataset is looking at how is likely to default on their credit. The following steps will be conducted in this analysis.
- Load the libraries and dataset
- Deal with missing data
- Some descriptive stats
- Normality check
- Model development
This is not an exhaustive analysis but rather a simple one for demonstration purposes. The dataset is available here
Load Libraries and Data
Here are some packages we will need
import pandas as pd import matplotlib.pyplot as plt import seaborn as sns from scipy.stats import norm from sklearn import tree from scipy import stats from sklearn import metrics
You can load the data with the code below
df_train=pd.read_csv('/application_train.csv')
You can examine what variables are available with the code below. This is not displayed here because it is rather long
df_train.columns df_train.head()
Missing Data
I prefer to deal with missing data first because missing values can cause errors throughout the analysis if they are not dealt with immediately. The code below calculates the percentage of missing data in each column.
total=df_train.isnull().sum().sort_values(ascending=False)
percent=(df_train.isnull().sum()/df_train.isnull().count()).sort_values(ascending=False)
missing_data=pd.concat([total,percent],axis=1,keys=['Total','Percent'])
missing_data.head()
Total Percent
COMMONAREA_MEDI 214865 0.698723
COMMONAREA_AVG 214865 0.698723
COMMONAREA_MODE 214865 0.698723
NONLIVINGAPARTMENTS_MODE 213514 0.694330
NONLIVINGAPARTMENTS_MEDI 213514 0.694330
Only the first five values are printed. You can see that some variables have a large amount of missing data. As such, they are probably worthless for inclusion in additional analysis. The code below removes all variables with any missing data.
pct_null = df_train.isnull().sum() / len(df_train) missing_features = pct_null[pct_null > 0.0].index df_train.drop(missing_features, axis=1, inplace=True)
You can use the .head() function if you want to see how many variables are left.
Data Description & Visualization
For demonstration purposes, we will print descriptive stats and make visualizations of a few of the variables that are remaining.
round(df_train['AMT_CREDIT'].describe()) Out[8]: count 307511.0 mean 599026.0 std 402491.0 min 45000.0 25% 270000.0 50% 513531.0 75% 808650.0 max 4050000.0 sns.distplot(df_train['AMT_CREDIT']

round(df_train['AMT_INCOME_TOTAL'].describe()) Out[10]: count 307511.0 mean 168798.0 std 237123.0 min 25650.0 25% 112500.0 50% 147150.0 75% 202500.0 max 117000000.0 sns.distplot(df_train['AMT_INCOME_TOTAL']

I think you are getting the point. You can also look at categorical variables using the groupby() function.
We also need to address categorical variables in terms of creating dummy variables. This is so that we can develop a model in the future. Below is the code for dealing with all the categorical variables and converting them to dummy variable’s
df_train.groupby('NAME_CONTRACT_TYPE').count()
dummy=pd.get_dummies(df_train['NAME_CONTRACT_TYPE'])
df_train=pd.concat([df_train,dummy],axis=1)
df_train=df_train.drop(['NAME_CONTRACT_TYPE'],axis=1)
df_train.groupby('CODE_GENDER').count()
dummy=pd.get_dummies(df_train['CODE_GENDER'])
df_train=pd.concat([df_train,dummy],axis=1)
df_train=df_train.drop(['CODE_GENDER'],axis=1)
df_train.groupby('FLAG_OWN_CAR').count()
dummy=pd.get_dummies(df_train['FLAG_OWN_CAR'])
df_train=pd.concat([df_train,dummy],axis=1)
df_train=df_train.drop(['FLAG_OWN_CAR'],axis=1)
df_train.groupby('FLAG_OWN_REALTY').count()
dummy=pd.get_dummies(df_train['FLAG_OWN_REALTY'])
df_train=pd.concat([df_train,dummy],axis=1)
df_train=df_train.drop(['FLAG_OWN_REALTY'],axis=1)
df_train.groupby('NAME_INCOME_TYPE').count()
dummy=pd.get_dummies(df_train['NAME_INCOME_TYPE'])
df_train=pd.concat([df_train,dummy],axis=1)
df_train=df_train.drop(['NAME_INCOME_TYPE'],axis=1)
df_train.groupby('NAME_EDUCATION_TYPE').count()
dummy=pd.get_dummies(df_train['NAME_EDUCATION_TYPE'])
df_train=pd.concat([df_train,dummy],axis=1)
df_train=df_train.drop(['NAME_EDUCATION_TYPE'],axis=1)
df_train.groupby('NAME_FAMILY_STATUS').count()
dummy=pd.get_dummies(df_train['NAME_FAMILY_STATUS'])
df_train=pd.concat([df_train,dummy],axis=1)
df_train=df_train.drop(['NAME_FAMILY_STATUS'],axis=1)
df_train.groupby('NAME_HOUSING_TYPE').count()
dummy=pd.get_dummies(df_train['NAME_HOUSING_TYPE'])
df_train=pd.concat([df_train,dummy],axis=1)
df_train=df_train.drop(['NAME_HOUSING_TYPE'],axis=1)
df_train.groupby('ORGANIZATION_TYPE').count()
dummy=pd.get_dummies(df_train['ORGANIZATION_TYPE'])
df_train=pd.concat([df_train,dummy],axis=1)
df_train=df_train.drop(['ORGANIZATION_TYPE'],axis=1)
You have to be careful with this because now you have many variables that are not necessary. For every categorical variable you must remove at least one category in order for the model to work properly. Below we did this manually.
df_train=df_train.drop(['Revolving loans','F','XNA','N','Y','SK_ID_CURR,''Student','Emergency','Lower secondary','Civil marriage','Municipal apartment'],axis=1)
Below are some boxplots with the target variable and other variables in the dataset.
f,ax=plt.subplots(figsize=(8,6)) fig=sns.boxplot(x=df_train['TARGET'],y=df_train['AMT_INCOME_TOTAL'])

There is a clear outlier there. Below is another boxplot with a different variable
f,ax=plt.subplots(figsize=(8,6)) fig=sns.boxplot(x=df_train['TARGET'],y=df_train['CNT_CHILDREN'])

It appears several people have more than 10 children. This is probably a typo.
Below is a correlation matrix using a heatmap technique
corrmat=df_train.corr() f,ax=plt.subplots(figsize=(12,9)) sns.heatmap(corrmat,vmax=.8,square=True)

The heatmap is nice but it is hard to really appreciate what is happening. The code below will sort the correlations from least to strongest, so we can remove high correlations.
c = df_train.corr().abs() s = c.unstack() so = s.sort_values(kind="quicksort") print(so.head()) FLAG_DOCUMENT_12 FLAG_MOBIL 0.000005 FLAG_MOBIL FLAG_DOCUMENT_12 0.000005 Unknown FLAG_MOBIL 0.000005 FLAG_MOBIL Unknown 0.000005 Cash loans FLAG_DOCUMENT_14 0.000005
The list is to long to show here but the following variables were removed for having a high correlation with other variables.
df_train=df_train.drop(['WEEKDAY_APPR_PROCESS_START','FLAG_EMP_PHONE','REG_CITY_NOT_WORK_CITY','REGION_RATING_CLIENT','REG_REGION_NOT_WORK_REGION'],axis=1)
Below we check a few variables for homoscedasticity, linearity, and normality using plots and histograms
sns.distplot(df_train['AMT_INCOME_TOTAL'],fit=norm) fig=plt.figure() res=stats.probplot(df_train['AMT_INCOME_TOTAL'],plot=plt)


This is not normal
sns.distplot(df_train['AMT_CREDIT'],fit=norm) fig=plt.figure() res=stats.probplot(df_train['AMT_CREDIT'],plot=plt)


This is not normal either. We could do transformations, or we can make a non-linear model instead.
Model Development
Now comes the easy part. We will make a decision tree using only some variables to predict the target. In the code below we make are X and y dataset.
X=df_train[['Cash loans','DAYS_EMPLOYED','AMT_CREDIT','AMT_INCOME_TOTAL','CNT_CHILDREN','REGION_POPULATION_RELATIVE']] y=df_train['TARGET']
The code below fits are model and makes the predictions
clf=tree.DecisionTreeClassifier(min_samples_split=20) clf=clf.fit(X,y) y_pred=clf.predict(X)
Below is the confusion matrix followed by the accuracy
print (pd.crosstab(y_pred,df_train['TARGET'])) TARGET 0 1 row_0 0 280873 18493 1 1813 6332 accuracy_score(y_pred,df_train['TARGET']) Out[47]: 0.933966589813047
Lastly, we can look at the precision, recall, and f1 score
print(metrics.classification_report(y_pred,df_train['TARGET']))
precision recall f1-score support
0 0.99 0.94 0.97 299366
1 0.26 0.78 0.38 8145
micro avg 0.93 0.93 0.93 307511
macro avg 0.62 0.86 0.67 307511
weighted avg 0.97 0.93 0.95 307511
This model looks rather good in terms of accuracy of the training set. It actually impressive that we could use so few variables from such a large dataset and achieve such a high degree of accuracy.
Conclusion
Data exploration and analysis is the primary task of a data scientist. This post was just an example of how this can be approached. Of course, there are many other creative ways to do this but the simplistic nature of this analysis yielded strong results
Confidence Intervals for Proportions VIDEO
Calculating Confidence Intervals for Proportions

Hierarchical Regression in R
In this post, we will learn how to conduct a hierarchical regression analysis in R. Hierarchical regression analysis is used in situation in which you want to see if adding additional variables to your model will significantly change the r2 when accounting for the other variables in the model. This approach is a model comparison approach and not necessarily a statistical one.
We are going to use the “Carseats” dataset from the ISLR package. Our goal will be to predict total sales using the following independent variables in three different models.
model 1 = intercept only
model 2 = Sales~Urban + US + ShelveLoc
model 3 = Sales~Urban + US + ShelveLoc + price + income
model 4 = Sales~Urban + US + ShelveLoc + price + income + Advertising
Often the primary goal with hierarchical regression is to show that the addition of a new variable builds or improves upon a previous model in a statistically significant way. For example, if a previous model was able to predict the total sales of an object using three variables you may want to see if a new additional variable you have in mind may improve model performance. Another way to see this is in the following research question
Is a model that explains the total sales of an object with Urban location, US location, shelf location, price, income and advertising cost as independent variables superior in terms of R2 compared to a model that explains total sales with Urban location, US location, shelf location, price and income as independent variables?
In this complex research question we essentially want to know if adding advertising cost will improve the model significantly in terms of the r square. The formal steps that we will following to complete this analysis is as follows.
- Build sequential (nested) regression models by adding variables at each step.
- Run ANOVAs in order to compute the R2
- Compute difference in sum of squares for each step
- Check F-statistics and p-values for the SS differences.
- Compare sum of squares between models from ANOVA results.
- Compute increase in R2 from sum of square difference
- Run regression to obtain the coefficients for each independent variable.
We will now begin our analysis. Below is some initial code
library(ISLR)
data("Carseats")Model Development
We now need to create our models. Model 1 will not have any variables in it and will be created for the purpose of obtaining the total sum of squares. Model 2 will include demographic variables. Model 3 will contain the initial model with the continuous independent variables. Lastly, model 4 will contain all the information of the previous models with the addition of the continuous independent variable of advertising cost. Below is the code.
model1 = lm(Sales~1,Carseats)
model2=lm(Sales~Urban + US + ShelveLoc,Carseats)
model3=lm(Sales~Urban + US + ShelveLoc + Price + Income,Carseats)
model4=lm(Sales~Urban + US + ShelveLoc + Price + Income + Advertising,Carseats)We can now turn to the ANOVA analysis for model comparison #ANOVA Calculation We will use the anova() function to calculate the total sum of square for model 0. This will serve as a baseline for the other models for calculating r square
anova(model1,model2,model3,model4)## Analysis of Variance Table
##
## Model 1: Sales ~ 1
## Model 2: Sales ~ Urban + US + ShelveLoc
## Model 3: Sales ~ Urban + US + ShelveLoc + Price + Income
## Model 4: Sales ~ Urban + US + ShelveLoc + Price + Income + Advertising
## Res.Df RSS Df Sum of Sq F Pr(>F)
## 1 399 3182.3
## 2 395 2105.4 4 1076.89 89.165 < 2.2e-16 ***
## 3 393 1299.6 2 805.83 133.443 < 2.2e-16 ***
## 4 392 1183.6 1 115.96 38.406 1.456e-09 ***
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1For now, we are only focusing on the residual sum of squares. Here is a basic summary of what we know as we compare the models.
model 1 = sum of squares = 3182.3
model 2 = sum of squares = 2105.4 (with demographic variables of Urban, US, and ShelveLoc)
model 3 = sum of squares = 1299.6 (add price and income)
model 4 = sum of squares = 1183.6 (add Advertising)
Each model is statistical significant which means adding each variable lead to some improvement.
By adding price and income to the model we were able to improve the model in a statistically significant way. The r squared increased by .25 below is how this was calculated.
2105.4-1299.6 #SS of Model 2 - Model 3## [1] 805.8805.8/ 3182.3 #SS difference of Model 2 and Model 3 divided by total sum of sqaure ie model 1## [1] 0.2532131When we add Advertising to the model the r square increases by .03. The calculation is below
1299.6-1183.6 #SS of Model 3 - Model 4## [1] 116116/ 3182.3 #SS difference of Model 3 and Model 4 divided by total sum of sqaure ie model 1## [1] 0.03645162Coefficients and R Square
We will now look at a summary of each model using the summary() function.
summary(model2)##
## Call:
## lm(formula = Sales ~ Urban + US + ShelveLoc, data = Carseats)
##
## Residuals:
## Min 1Q Median 3Q Max
## -6.713 -1.634 -0.019 1.738 5.823
##
## Coefficients:
## Estimate Std. Error t value Pr(>|t|)
## (Intercept) 4.8966 0.3398 14.411 < 2e-16 ***
## UrbanYes 0.0999 0.2543 0.393 0.6947
## USYes 0.8506 0.2424 3.510 0.0005 ***
## ShelveLocGood 4.6400 0.3453 13.438 < 2e-16 ***
## ShelveLocMedium 1.8168 0.2834 6.410 4.14e-10 ***
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
##
## Residual standard error: 2.309 on 395 degrees of freedom
## Multiple R-squared: 0.3384, Adjusted R-squared: 0.3317
## F-statistic: 50.51 on 4 and 395 DF, p-value: < 2.2e-16summary(model3)##
## Call:
## lm(formula = Sales ~ Urban + US + ShelveLoc + Price + Income,
## data = Carseats)
##
## Residuals:
## Min 1Q Median 3Q Max
## -4.9096 -1.2405 -0.0384 1.2754 4.7041
##
## Coefficients:
## Estimate Std. Error t value Pr(>|t|)
## (Intercept) 10.280690 0.561822 18.299 < 2e-16 ***
## UrbanYes 0.219106 0.200627 1.092 0.275
## USYes 0.928980 0.191956 4.840 1.87e-06 ***
## ShelveLocGood 4.911033 0.272685 18.010 < 2e-16 ***
## ShelveLocMedium 1.974874 0.223807 8.824 < 2e-16 ***
## Price -0.057059 0.003868 -14.752 < 2e-16 ***
## Income 0.013753 0.003282 4.190 3.44e-05 ***
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
##
## Residual standard error: 1.818 on 393 degrees of freedom
## Multiple R-squared: 0.5916, Adjusted R-squared: 0.5854
## F-statistic: 94.89 on 6 and 393 DF, p-value: < 2.2e-16summary(model4)##
## Call:
## lm(formula = Sales ~ Urban + US + ShelveLoc + Price + Income +
## Advertising, data = Carseats)
##
## Residuals:
## Min 1Q Median 3Q Max
## -5.2199 -1.1703 0.0225 1.0826 4.1124
##
## Coefficients:
## Estimate Std. Error t value Pr(>|t|)
## (Intercept) 10.299180 0.536862 19.184 < 2e-16 ***
## UrbanYes 0.198846 0.191739 1.037 0.300
## USYes -0.128868 0.250564 -0.514 0.607
## ShelveLocGood 4.859041 0.260701 18.638 < 2e-16 ***
## ShelveLocMedium 1.906622 0.214144 8.903 < 2e-16 ***
## Price -0.057163 0.003696 -15.467 < 2e-16 ***
## Income 0.013750 0.003136 4.384 1.50e-05 ***
## Advertising 0.111351 0.017968 6.197 1.46e-09 ***
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
##
## Residual standard error: 1.738 on 392 degrees of freedom
## Multiple R-squared: 0.6281, Adjusted R-squared: 0.6214
## F-statistic: 94.56 on 7 and 392 DF, p-value: < 2.2e-16You can see for yourself the change in the r square. From model 2 to model 3 there is a 26 point increase in r square just as we calculated manually. From model 3 to model 4 there is a 3 point increase in r square. The purpose of the anova() analysis was determined if the significance of the change meet a statistical criterion, The lm() function reports a change but not the significance of it.
Conclusion
Hierarchical regression is just another potential tool for the statistical researcher. It provides you with a way to develop several models and compare the results based on any potential improvement in the r square.
Confidence Intervals VIDEO
Calculating Confidence intervals
Standard Deviation VIDEO
Calculating standard deviation

Elastic Net Regression in Python

Elastic net regression combines the power of ridge and lasso regression into one algorithm. What this means is that with elastic net the algorithm can remove weak variables altogether as with lasso or to reduce them to close to zero as with ridge. All of these algorithms are examples of regularized regression.
This post will provide an example of elastic net regression in Python. Below are the steps of the analysis.
- Data preparation
- Baseline model development
- Elastic net model development
To accomplish this, we will use the Fair dataset from the pydataset library. Our goal will be to predict marriage satisfaction based on the other independent variables. Below is some initial code to begin the analysis.
from pydataset import data
import numpy as np
import pandas as pd
pd.set_option('display.max_rows', 5000)
pd.set_option('display.max_columns', 5000)
pd.set_option('display.width', 10000)
from sklearn.model_selection import GridSearchCV
from sklearn.linear_model import ElasticNet
from sklearn.linear_model import LinearRegression
from sklearn.metrics import mean_squared_error
Data Preparation
We will now load our data. The only preparation that we need to do is convert the factor variables to dummy variables. Then we will make our and y datasets. Below is the code.
df=pd.DataFrame(data('Fair'))
df.loc[df.sex== 'male', 'sex'] = 0
df.loc[df.sex== 'female','sex'] = 1
df['sex'] = df['sex'].astype(int)
df.loc[df.child== 'no', 'child'] = 0
df.loc[df.child== 'yes','child'] = 1
df['child'] = df['child'].astype(int)
X=df[['religious','age','sex','ym','education','occupation','nbaffairs']]
y=df['rate']
We can now proceed to creating the baseline model
Baseline Model
This model is a basic regression model for the purpose of comparison. We will instantiate our regression model, use the fit command and finally calculate the mean squared error of the data. The code is below.
regression=LinearRegression()
regression.fit(X,y)
first_model=(mean_squared_error(y_true=y,y_pred=regression.predict(X)))
print(first_model)
1.0498738644696668
This mean standard error score of 1.05 is our benchmark for determining if the elastic net model will be better or worst. Below are the coefficients of this first model. We use a for loop to go through the model and the zip function to combine the two columns.
coef_dict_baseline = {}
for coef, feat in zip(regression.coef_,X.columns):
coef_dict_baseline[feat] = coef
coef_dict_baseline
Out[63]:
{'religious': 0.04235281110639178,
'age': -0.009059645428673819,
'sex': 0.08882013337087094,
'ym': -0.030458802565476516,
'education': 0.06810255742293699,
'occupation': -0.005979506852998164,
'nbaffairs': -0.07882571247653956}
We will now move to making the elastic net model.
Elastic Net Model
Elastic net, just like ridge and lasso regression, requires normalize data. This argument is set inside the ElasticNet function. The second thing we need to do is create our grid. This is the same grid as we create for ridge and lasso in prior posts. The only thing that is new is the l1_ratio argument.
When the l1_ratio is set to 0 it is the same as ridge regression. When l1_ratio is set to 1 it is lasso. Elastic net is somewhere between 0 and 1 when setting the l1_ratio. Therefore, in our grid, we need to set several values of this argument. Below is the code.
elastic=ElasticNet(normalize=True)
search=GridSearchCV(estimator=elastic,param_grid={'alpha':np.logspace(-5,2,8),'l1_ratio':[.2,.4,.6,.8]},scoring='neg_mean_squared_error',n_jobs=1,refit=True,cv=10)
We will now fit our model and display the best parameters and the best results we can get with that setup.
search.fit(X,y)
search.best_params_
Out[73]: {'alpha': 0.001, 'l1_ratio': 0.8}
abs(search.best_score_)
Out[74]: 1.0816514028705004
The best hyperparameters was an alpha set to 0.001 and a l1_ratio of 0.8. With these settings we got an MSE of 1.08. This is above our baseline model of MSE 1.05 for the baseline model. Which means that elastic net is doing worse than linear regression. For clarity, we will set our hyperparameters to the recommended values and run on the data.
elastic=ElasticNet(normalize=True,alpha=0.001,l1_ratio=0.75)
elastic.fit(X,y)
second_model=(mean_squared_error(y_true=y,y_pred=elastic.predict(X)))
print(second_model)
1.0566430678343806
Now our values are about the same. Below are the coefficients
coef_dict_baseline = {}
for coef, feat in zip(elastic.coef_,X.columns):
coef_dict_baseline[feat] = coef
coef_dict_baseline
Out[76]:
{'religious': 0.01947541724957858,
'age': -0.008630896492807691,
'sex': 0.018116464568090795,
'ym': -0.024224831274512956,
'education': 0.04429085595448633,
'occupation': -0.0,
'nbaffairs': -0.06679513627963515}
The coefficients are mostly the same. Notice that occupation was completely removed from the model in the elastic net version. This means that this values was no good to the algorithm. Traditional regression cannot do this.
Conclusion
This post provided an example of elastic net regression. Elastic net regression allows for the maximum flexibility in terms of finding the best combination of ridge and lasso regression characteristics. This flexibility is what gives elastic net its power.

Lasso Regression with Python
Lasso regression is another form of regularized regression. With this particular version, the coefficient of a variable can be reduced all the way to zero through the use of the l1 regularization. This is in contrast to ridge regression which never completely removes a variable from an equation as it employs l2 regularization.

Regularization helps to stabilize estimates as well as deal with bias and variance in a model. In this post, we will use the “CaSchools” dataset from the pydataset library. Our goal will be to predict test scores based on several independent variables. The steps we will follow are as follows.
- Data preparation
- Develop a baseline linear model
- Develop lasso regression model
The initial code is as follows
from pydataset import data
import numpy as np
import pandas as pd
from sklearn.linear_model import LinearRegression
from sklearn.metrics import mean_squared_error
from sklearn.model_selection import GridSearchCV
from sklearn.linear_model import Lasso
df=pd.DataFrame(data(‘Caschool’))
Data Preparation
The data preparation is simple in this example. We only have to store the desired variables in our X and y datasets. We are not using all of the variables. Some were left out because they were highly correlated. Lasso is able to deal with this to a certain extent w=but it was decided to leave them out anyway. Below is the code.
X=df[['teachers','calwpct','mealpct','compstu','expnstu','str','avginc','elpct']]
y=df['testscr']
Baseline Model
We can now run our baseline model. This will give us a measure of comparison for the lasso model. Our metric is the mean squared error. Below is the code with the results of the model.
regression=LinearRegression()
regression.fit(X,y)
LinearRegression(copy_X=True, fit_intercept=True, n_jobs=1, normalize=False)
first_model=(mean_squared_error(y_true=y,y_pred=regression.predict(X)))
print(first_model)
69.07380530137416
First, we instantiate the LinearRegression class. Then, we run the .fit method to do the analysis. Next, we predicted future values of our regression model and save the results to the object first_model. Lastly, we printed the results.
Below are the coefficient for the baseline regression model.
coef_dict_baseline = {}
for coef, feat in zip(regression.coef_,X.columns):
coef_dict_baseline[feat] = coef
coef_dict_baseline
Out[52]:
{'teachers': 0.00010011947964873427,
'calwpct': -0.07813766458116565,
'mealpct': -0.3754719080127311,
'compstu': 11.914006268826652,
'expnstu': 0.001525630709965126,
'str': -0.19234209691788984,
'avginc': 0.6211690806021222,
'elpct': -0.19857026121348267}
The for loop simply combines the features in our model with their coefficients. With this information we can now make our lasso model and compare the results.
Lasso Model
For our lasso model, we have to determine what value to set the l1 or alpha to prior to creating the model. This can be done with the grid function, This function allows you to assess several models with different l1 settings. Then python will tell which setting is the best. Below is the code.
lasso=Lasso(normalize=True)
search=GridSearchCV(estimator=lasso,param_grid={'alpha':np.logspace(-5,2,8)},scoring='neg_mean_squared_error',n_jobs=1,refit=True,cv=10)
search.fit(X,y)
We start be instantiate lasso with normalization set to true. It is important to scale data when doing regularized regression. Next, we setup our grid, we include the estimator, and parameter grid, and scoring. The alpha is set using logspace. We want values between -5 and 2, and we want 8 evenly spaced settings for the alpha. The other arguments include cv which stands for cross-validation. n_jobs effects processing and refit updates the parameters.
After completing this, we used the fit function. The code below indicates the appropriate alpha and the expected score if we ran the model with this alpha setting.
search.best_params_
Out[55]: {'alpha': 1e-05}
abs(search.best_score_)
Out[56]: 85.38831122904011
`The alpha is set almost to zero, which is the same as a regression model. You can also see that the mean squared error is actually worse than in the baseline model. In the code below, we run the lasso model with the recommended alpha setting and print the results.
lasso=Lasso(normalize=True,alpha=1e-05)
lasso.fit(X,y)
second_model=(mean_squared_error(y_true=y,y_pred=lasso.predict(X)))
print(second_model)
69.0738055527604
The value for the second model is almost the same as the first one. The tiny difference is due to the fact that there is some penalty involved. Below are the coefficient values.
coef_dict_baseline = {}
for coef, feat in zip(lasso.coef_,X.columns):
coef_dict_baseline[feat] = coef
coef_dict_baseline
Out[63]:
{'teachers': 9.795933425676567e-05,
'calwpct': -0.07810938255735576,
'mealpct': -0.37548182158171706,
'compstu': 11.912164626067028,
'expnstu': 0.001525439984250718,
'str': -0.19225486069458508,
'avginc': 0.6211695477945162,
'elpct': -0.1985510490295491}
The coefficient values are also slightly different. The only difference is the teachers variable was essentially set to zero. This means that it is not a useful variable for predicting testscrs. That is ironic to say the least.
Conclusion
Lasso regression is able to remove variables that are not adequate predictors of the outcome variable. Doing this in Python is fairly simple. This yet another tool that can be used in statistical analysis.

Ridge Regression in Python

Ridge regression is one of several regularized linear models. Regularization is the process of penalizing coefficients of variables either by removing them and or reduce their impact. Ridge regression reduces the effect of problematic variables close to zero but never fully removes them.
We will go through an example of ridge regression using the VietNamI dataset available in the pydataset library. Our goal will be to predict expenses based on the variables available. We will complete this task using the following steps/
- Data preparation
- Baseline model development
- Ridge regression model
Below is the initial code
from pydataset import data
import numpy as np
import pandas as pd
from sklearn.model_selection import GridSearchCV
from sklearn.linear_model import Ridge
from sklearn.linear_model import LinearRegression
from sklearn.metrics import mean_squared_erro
Data Preparation
The data preparation is simple. All we have to do is load the data and convert the sex variable to a dummy variable. We also need to set up our X and y datasets. Below is the code.
df=pd.DataFrame(data('VietNamI'))
df.loc[df.sex== 'male', 'sex'] = 0
df.loc[df.sex== 'female','sex'] = 1
df['sex'] = df['sex'].astype(int)
X=df[['pharvis','age','sex','married','educ','illness','injury','illdays','actdays','insurance']]
y=df['lnhhexp'
We can now create our baseline regression model.
Baseline Model
The metric we are using is the mean squared error. Below is the code and output for our baseline regression model. This is a model that has no regularization to it. Below is the code.
regression=LinearRegression()
regression.fit(X,y)
first_model=(mean_squared_error(y_true=y,y_pred=regression.predict(X)))
print(first_model)
0.35528915032173053
This value of 0.355289 will be our indicator to determine if the regularized ridge regression model is superior or not.
Ridge Model
In order to create our ridge model we need to first determine the most appropriate value for the l2 regularization. L2 is the name of the hyperparameter that is used in ridge regression. Determining the value of a hyperparameter requires the use of a grid. In the code below, we first are ridge model and indicate normalization in order to get better estimates. Next we setup the grid that we will use. Below is the code.
ridge=Ridge(normalize=True)
search=GridSearchCV(estimator=ridge,param_grid={'alpha':np.logspace(-5,2,8)},scoring='neg_mean_squared_error',n_jobs=1,refit=True,cv=10)
The search object has several arguments within it. Alpha is hyperparameter we are trying to set. The log space is the range of values we want to test. We want the log of -5 to 2, but we only get 8 values from within that range evenly spread out. Are metric is the mean squared error. Refit set true means to adjust the parameters while modeling and cv is the number of folds to develop for the cross-validation. We can now use the .fit function to run the model and then use the .best_params_ and .best_scores_ function to determine the model;s strength. Below is the code.
search.fit(X,y)
search.best_params_
{'alpha': 0.01}
abs(search.best_score_)
0.3801489007094425
The best_params_ tells us what to set alpha too which in this case is 0.01. The best_score_ tells us what the best possible mean squared error is. In this case, the value of 0.38 is worse than what the baseline model was. We can confirm this by fitting our model with the ridge information and finding the mean squared error. This is done below.
ridge=Ridge(normalize=True,alpha=0.01)
ridge.fit(X,y)
second_model=(mean_squared_error(y_true=y,y_pred=ridge.predict(X)))
print(second_model)
0.35529321992606566
The 0.35 is lower than the 0.38. This is because the last results are not cross-validated. In addition, these results indicate that there is little difference between the ridge and baseline models. This is confirmed with the coefficients of each model found below.
coef_dict_baseline = {}
for coef, feat in zip(regression.coef_,data("VietNamI").columns):
coef_dict_baseline[feat] = coef
coef_dict_baseline
Out[188]:
{'pharvis': 0.013282050886950674,
'lnhhexp': 0.06480086550467873,
'age': 0.004012412278795848,
'sex': -0.08739614349708981,
'married': 0.075276463838362,
'educ': -0.06180921300600292,
'illness': 0.040870384578962596,
'injury': -0.002763768716569026,
'illdays': -0.006717063310893158,
'actdays': 0.1468784364977112}
coef_dict_ridge = {}
for coef, feat in zip(ridge.coef_,data("VietNamI").columns):
coef_dict_ridge[feat] = coef
coef_dict_ridge
Out[190]:
{'pharvis': 0.012881937698185289,
'lnhhexp': 0.06335455237380987,
'age': 0.003896623321297935,
'sex': -0.0846541637961565,
'married': 0.07451889604357693,
'educ': -0.06098723778992694,
'illness': 0.039430607922053884,
'injury': -0.002779341753010467,
'illdays': -0.006551280792122459,
'actdays': 0.14663287713359757}
The coefficient values are about the same. This means that the penalization made little difference with this dataset.
Conclusion
Ridge regression allows you to penalize variables based on their useful in developing the model. With this form of regularized regression the coefficients of the variables is never set to zero. Other forms of regularization regression allows for the total removal of variables. One example of this is lasso regression.

Random Forest Regression in Python
Random forest is simply the making of dozens if not thousands of decision trees. The decision each tree makes about an example are then tallied for the purpose of voting with the classification that receives the most votes winning. For regression, the results of the trees are averaged in order to give the most accurate results
In this post, we will use the cancer dataset from the pydataset module to predict the age of people. Below is some initial code.
import pandas as pd import numpy as np from pydataset import data from sklearn.model_selection import train_test_split from sklearn.model_selection import cross_val_score from sklearn.ensemble import RandomForestRegressor from sklearn.metrics import mean_squared_error
We can load our dataset as df, drop all NAs, and create our dataset that contains the independent variables and a separate dataset that includes the dependent variable of age. The code is below
df = data('cancer')
df=df.dropna()
X=df[['time','status',"sex","ph.ecog",'ph.karno','pat.karno','meal.cal','wt.loss']]
y=df['age']
Next, we need to set up our train and test sets using a 70/30 split. After that, we set up our model using the RandomForestRegressor function. n_estimators is the number of trees we want to create and the random_state argument is for supporting reproducibility. The code is below
x_train, x_test, y_train, y_test = train_test_split(X, y, test_size=0.3, random_state=0) h=RandomForestRegressor(n_estimators=100,random_state=1)
We can now run our model and test it. Running the model requires the .fit() function and testing involves the .predict() function. The results of the test are found using the mean_squared_error() function.
h.fit(x_train,y_train) y_pred=h.predict(x_test) mean_squared_error(y_test,y_pred) 71.75780196078432
The MSE of 71.75 is only useful for model comparison and has little meaning by its self. Another way to assess the model is by determining variable importance. This helps you to determine in a descriptive way the strongest variables for the regression model. The code is below followed by the plot of the variables.
model_ranks=pd.Series(h.feature_importances_,index=x_train.columns,name="Importance").sort_values(ascending=True,inplace=False) ax=model_ranks.plot(kind='barh')

As you can see, the strongest predictors of age include calories per meal, weight loss, and time sick. Sex and whether the person is censored or dead make a smaller difference. This makes sense as younger people eat more and probably lose more weight because they are heavier initially when dealing with cancer.
Conclusison
This post provided an example of the use of regression with random forest. Through the use of ensemble voting, you can improve the accuracy of your models. This is a distinct power that is not available with other machine learning algorithm.

Support Vector Machines Regression with Python
This post will provide an example of how to do regression with support vector machines SVM. SVM is a complex algorithm that allows for the development of non-linear models. This is particularly useful for messy data that does not have clear boundaries.
The steps that we will use are listed below
- Data preparation
- Model Development
We will use two different kernels in our analysis. The LinearSVR kernel and SVR kernel. The difference between these two kernels has to do with slight changes in the calculations of the boundaries between classes.
Data Preparation
We are going to use the OFP dataset available in the pydataset module. This dataset was used previously for classification with SVM on this site. Our plan this time is that we want to predict family inc (famlinc), which is a continuous variable. Below is some initial code.
import numpy as np import pandas as pd from pydataset import data from sklearn import svm from sklearn import model_selection from statsmodels.tools.eval_measures import mse
We now need to load our dataset and remove any missing values.
df=pd.DataFrame(data('OFP'))
df=df.dropna()
AS in the previous post, we need to change the text variables into dummy variables and we also need to scale the data. The code below creates the dummy variables, removes variables that are not needed, and also scales the data.
dummy=pd.get_dummies(df['black'])
df=pd.concat([df,dummy],axis=1)
df=df.rename(index=str, columns={"yes": "black_person"})
df=df.drop('no', axis=1)
dummy=pd.get_dummies(df['sex'])
df=pd.concat([df,dummy],axis=1)
df=df.rename(index=str, columns={"male": "Male"})
df=df.drop('female', axis=1)
dummy=pd.get_dummies(df['employed'])
df=pd.concat([df,dummy],axis=1)
df=df.rename(index=str, columns={"yes": "job"})
df=df.drop('no', axis=1)
dummy=pd.get_dummies(df['maried'])
df=pd.concat([df,dummy],axis=1)
df=df.rename(index=str, columns={"no": "single"})
df=df.drop('yes', axis=1)
dummy=pd.get_dummies(df['privins'])
df=pd.concat([df,dummy],axis=1)
df=df.rename(index=str, columns={"yes": "insured"})
df=df.drop('no', axis=1)
df=df.drop(['black','sex','maried','employed','privins','medicaid','region','hlth'],axis=1)
df = (df - df.min()) / (df.max() - df.min())
df.head()

We now need to set up our datasets. The X dataset will contain the independent variables while the y dataset will contain the dependent variable
X=df[['ofp','ofnp','opp','opnp','emr','hosp','numchron','adldiff','age','school','single','black_person','Male','job','insured']] y=df['faminc']
We can now move to model development
Model Development
We now need to create our train and test sets for or X and y datasets. We will do a 70/30 split of the data. Below is the code
X_train,X_test,y_train,y_test=model_selection.train_test_split(X,y,test_size=.3,random_state=1)
Next, we will create our two models with the code below.
h1=svm.SVR() h2=svm.LinearSVR()
We will now run our first model and assess the results. Our metric is the mean squared error. Generally, the lower the number the better. We will use the .fit() function to train the model and the .predict() function for test the model

The mse was 0.27. This number means nothing only and is only beneficial for comparison reasons. Therefore, the second model will be judged as better or worst only if the mse is lower than 0.27. Below are the results of the second model.

We can see that the mse for our second model is 0.34 which is greater than the mse for the first model. This indicates that the first model is superior based on the current results and parameter settings.
Conclusion
This post provided an example of how to use SVM for regression.

Multiple Regression in Python
In this post, we will go through the process of setting up and a regression model with a training and testing set using Python. We will use the insurance dataset from kaggle. Our goal will be to predict charges. In this analysis, the following steps will be performed.
- Data preparation
- Model training
- model testing
Data Preparation
Below is a list of the modules we will need in order to complete the analysis
import matplotlib.pyplot as plt import pandas as pd from sklearn import linear_model,model_selection, feature_selection,preprocessing import statsmodels.formula.api as sm from statsmodels.tools.eval_measures import mse from statsmodels.tools.tools import add_constant from sklearn.metrics import mean_squared_error
After you download the dataset you need to load it and take a look at it. You will use the .read_csv function from pandas to load the data and .head() function to look at the data. Below is the code and the output.
insure=pd.read_csv('YOUR LOCATION HERE')

We need to create some dummy variables for sex, smoker, and region. We will address that in a moment, right now we will look at descriptive stats for our continuous variables. We will use the .describe() function for descriptive stats and the .corr() function to find the correlations.

The descriptives are left for your own interpretation. As for the correlations, they are generally weak which is an indication that regression may be appropriate.
As mentioned earlier, we need to make dummy variables sex, smoker, and region in order to do the regression analysis. To complete this we need to do the following.
- Use the pd.get_dummies function from pandas to create the dummy
- Save the dummy variable in an object called ‘dummy’
- Use the pd.concat function to add our new dummy variable to our ‘insure’ dataset
- Repeat this three times
Below is the code for doing this
dummy=pd.get_dummies(insure['sex']) insure=pd.concat([insure,dummy],axis=1) dummy=pd.get_dummies(insure['smoker']) insure=pd.concat([insure,dummy],axis=1) dummy=pd.get_dummies(insure['region']) insure=pd.concat([insure,dummy],axis=1) insure.head()

The .get_dummies function requires the name of the dataframe and in the brackets the name of the variable to convert. The .concat function requires the name of the two datasets to combine as well the axis on which to perform it.
We now need to remove the original text variables from the dataset. In addition, we need to remove the y variable “charges” because this is the dependent variable.
y = insure.charges insure=insure.drop(['sex', 'smoker','region','charges'], axis=1)
We can now move to model development.
Model Training
Are train and test sets are model with the model_selection.trainin_test_split function. We will do an 80-20 split of the data. Below is the code.
X_train, X_test, y_train, y_test = model_selection.train_test_split(insure, y, test_size=0.2)
In this single line of code, we create a train and test set of our independent variables and our dependent variable.
We can not run our regression analysis. This requires the use of the .OLS function from statsmodels module. Below is the code.
answer=sm.OLS(y_train, add_constant(X_train)).fit()
In the code above inside the parentheses, we put the dependent variable(y_train) and the independent variables (X_train). However, we had to use the function add_constant to get the intercept for the output. All of this information is then used inside the .fit() function to fit a model.
To see the output you need to use the .summary() function as shown below.
answer.summary()

The assumption is that you know regression but our reading this post to learn python. Therefore, we will not go into great detail about the results. The r-square is strong, however, the region and gender are not statistically significant.
We will now move to model testing
Model Testing
Our goal here is to take the model that we developed and see how it does on other data. First, we need to predict values with the model we made with the new data. This is shown in the code below
ypred=answer.predict(add_constant(X_test))
We use the .predict() function for this action and we use the X_test data as well. With this information, we will calculate the mean squared error. This metric is useful for comparing models. We only made one model so it is not that useful in this situation. Below is the code and results.
print(mse(ypred,y_test)) 33678660.23480476
For our final trick, we will make a scatterplot with the predicted and actual values of the test set. In addition, we will calculate the correlation of the predict values and test set values. This is an alternative metric for assessing a model.

You can see the first two lines are for making the plot. Lines 3-4 are for making the correlation matrix and involves the .concat() function. The correlation is high at 0.86 which indicates the model is good at accurately predicting the values. THis is confirmed with the scatterplot which is almost a straight line.
Conclusion
IN this post we learned how to do a regression analysis in Python. We prepared the data, developed a model, and tested a model with an evaluation of it.

Logistic Regression in Python
This post will provide an example of a logistic regression analysis in Python. Logistic regression is commonly used when the dependent variable is categorical. Our goal will be to predict the gender of an example based on the other variables in the model. Below are the steps we will take to achieve this.
- Data preparation
- Model development
- Model testing
- Model evaluation
Data Preparation
The dataset we will use is the ‘Survey of Labour and Income Dynamics’ (SLID) dataset available in the pydataset module in Python. This dataset contains basic data on labor and income along with some demographic information. The initial code that we need is below.
import pandas as pd import statsmodels.api as sm import numpy as np import matplotlib.pyplot as plt from sklearn.linear_model import LogisticRegression from sklearn.model_selection import train_test_split from sklearn import metrics from pydataset import data
The code above loads all the modules and other tools we will need in this example. We now can load our data. In addition to loading the data, we will also look at the count and the characteristics of the variables. Below is the code.

At the top of this code, we create the ‘df’ object which contains our data from the “SLID”. Next, we used the .count() function to determine if there was any missing data and to see what variables were available. It appears that we have five variables and a lot of missing data as each variable has different amounts of data. Lastly, we used the .head() function to see what each variable contained. It appears that wages, education, and age are continuous variables well sex and language are categorical. The categorical variables will need to be converted to dummy variables as well.
The next thing we need to do is drop all the rows that are missing data since it is hard to create a model when data is missing. Below is the code and the output for this process.

In the code above, we used the .dropna() function to remove missing data. Then we used the .count() function to see how many rows remained. You can see that all the variables have the same number of rows which is important for model analysis. We will now make our dummy variables for sex and language in the code below.

Here is what we did,
- We used the .get_dummies function from pandas first on the sex variable. All this was stored in a new object called “dummy”
- We then combined the dummy and df datasets using the .concat() function. The axis =1 argument is for combing by column.
- We repeat steps 1 and 2 for the language variable
- Lastly, we used the .head() function to see the results
With this, we are ready to move to model development.
Model Development
The first thing we need to do is put all of the independent variables in one dataframe and the dependent variable in its own dataframe. Below is the code for this
X=df[['wages','education','age',"French","Other"]] y=df['Male']
Notice that we did not use every variable that was available. For the language variables, we only used “French” and “Other”. This is because when you make dummy variables you only need k-1 dummies created. Since the language variable had three categories we only need two dummy variables. Therefore, we excluded “English” because when “French” and “Other” are coded 0 it means that “English” is the characteristic of the example.
In addition, we only took “male” as our dependent variable because if “male” is set to 0 it means that example is female. We now need to create our train and test dataset. The code is below.
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3, random_state=0)
We created four datasets
- train dataset with the independent variables
- train dataset with the dependent variable
- test dataset with the independent variables
- test dataset with the independent variable
The split is 70/30 with 70% being used for the training and 30% being used for testing. This is the purpose of the “test_size” argument. we used the train_test_split function to do this. We can now run our model and get the results. Below is the code.

Here is what we did
- We used the .Logit() function from statsmodel to create the logistic model. Notice we used only the training data.
- We then use the .fit() function to get the results and stored this in the result object.
- Lastly, we printed the results in the ‘result’ object using the .summary()
There are some problems with the results. The Pseudo R-square is infinity which is usually. Also, you may have some error output about hessian inversion in your output. For these reasons, we cannot trust the results but will continue for the sake of learning.
The coefficients are clear. Only wage, education, and age are significant. In order to determine the probability you have to take the coefficient from the model and use the .exp() function from numpy. Below are the results.
np.exp(.08) Out[107]: 1.0832870676749586 np.exp(-0.06) Out[108]: 0.9417645335842487 np.exp(-.01) Out[109]: 0.9900498337491681
For the first value, for every unit wages increaser the probability that they are male increase 8%. For every 1 unit increase in education there probability of the person being male decrease 6%. Lastly, for every one unit increase in age the probability of the person being male decrease by 1%. Notice that we subtract 1 from the outputs to find the actual probability.
We will now move to model testing
Model Testing
To do this we first test our model with the code below
y_pred=result.predict(X_test)
We made the result object earlier. Now we just use the .predict() function with the X_test data. Next, we need to flag examples that the model believes has a 60% chance or greater of being male. The code is below
y_pred_flag=y_pred>.6
This creates a boolean object with True and False as the output. Now we will make our confusion matrix as well as other metrics for classification.

The results speak for themselves. There are a lot of false positives if you look at the confusion matrix. In addition precision, recall, and f1 are all low. Hopefully, the coding should be clear the main point is to be sure to use the test set dependent dataset (y_test) with the flag data you made in the previous step.
We will not make the ROC curve. For a strong model, it should have a strong elbow shape while with a weak model it will be a diagonal straight line.

The first plot is of our data. The second plot is what a really bad model would look like. As you can see there is littte difference between the two. Again this is because of all the false positives we have in the model. The actual coding should be clear. fpr is the false positive rate, tpr is the true positive rate. The function is .roc_curve. Inside goes the predict vs actual test data.
Conclusion
This post provided a demonstration of the use of logistic regression in Python. It is necessary to follow the steps above but keep in mind that this was a demonstration and the results are dubious.
Local Regression in R
Local regression uses something similar to nearest neighbor classification to generate a regression line. In local regression, nearby observations are used to fit the line rather than all observations. It is necessary to indicate the percentage of the observations you want R to use for fitting the local line. The name for this hyperparameter is the span. The higher the span the smoother the line becomes.
Local regression is great one there are only a handful of independent variables in the model. When the total number of variables becomes too numerous the model will struggle. As such, we will only fit a bivariate model. This will allow us to process the model and to visualize it.
In this post, we will use the “Clothing” dataset from the “Ecdat” package and we will examine innovation (inv2) relationship with total sales (tsales). Below is some initial code.
library(Ecdat)data(Clothing)
str(Clothing)## 'data.frame': 400 obs. of 13 variables:
## $ tsales : int 750000 1926395 1250000 694227 750000 400000 1300000 495340 1200000 495340 ...
## $ sales : num 4412 4281 4167 2670 15000 ...
## $ margin : num 41 39 40 40 44 41 39 28 41 37 ...
## $ nown : num 1 2 1 1 2 ...
## $ nfull : num 1 2 2 1 1.96 ...
## $ npart : num 1 3 2.22 1.28 1.28 ...
## $ naux : num 1.54 1.54 1.41 1.37 1.37 ...
## $ hoursw : int 76 192 114 100 104 72 161 80 158 87 ...
## $ hourspw: num 16.8 22.5 17.2 21.5 15.7 ...
## $ inv1 : num 17167 17167 292857 22207 22207 ...
## $ inv2 : num 27177 27177 71571 15000 10000 ...
## $ ssize : int 170 450 300 260 50 90 400 100 450 75 ...
## $ start : num 41 39 40 40 44 41 39 28 41 37 ...There is no data preparation in this example. The first thing we will do is fit two different models that have different values for the span hyperparameter. “fit” will have a span of .41 which means it will use 41% of the nearest examples. “fit2” will use .82. Below is the code.
fit<-loess(tsales~inv2,span = .41,data = Clothing)
fit2<-loess(tsales~inv2,span = .82,data = Clothing)In the code above, we used the “loess” function to fit the model. The “span” argument was set to .41 and .82.
We now need to prepare for the visualization. We begin by using the “range” function to find the distance from the lowest to the highest value. Then use the “seq” function to create a grid. Below is the code.
inv2lims<-range(Clothing$inv2)
inv2.grid<-seq(from=inv2lims[1],to=inv2lims[2])The information in the code above is for setting our x-axis in the plot. We are now ready to fit our model. We will fit the models and draw each regression line.
plot(Clothing$inv2,Clothing$tsales,xlim=inv2lims)
lines(inv2.grid,predict(fit,data.frame(inv2=inv2.grid)),col='blue',lwd=3)
lines(inv2.grid,predict(fit2,data.frame(inv2=inv2.grid)),col='red',lwd=3)
Not much difference in the two models. For our final task, we will predict with our “fit” model using all possible values of “inv2” and also fit the confidence interval lines.
pred<-predict(fit,newdata=inv2.grid,se=T)
plot(Clothing$inv2,Clothing$tsales)
lines(inv2.grid,pred$fit,col='red',lwd=3)
lines(inv2.grid,pred$fit+2*pred$se.fit,lty="dashed",lwd=2,col='blue')
lines(inv2.grid,pred$fit-2*pred$se.fit,lty="dashed",lwd=2,col='blue')
Conclusion
Local regression provides another way to model complex non-linear relationships in low dimensions. The example here provides just the basics of how this is done is much more complicated than described here.
Smoothing Splines in R
This post will provide information on smoothing splines. Smoothing splines are used in regression when we want to reduce the residual sum of squares by adding more flexibility to the regression line without allowing too much overfitting.
In order to do this, we must tune the parameter called the smoothing spline. The smoothing spline is essentially a natural cubic spline with a knot at every unique value of x in the model. Having this many knots can lead to severe overfitting. This is corrected for by controlling the degrees of freedom through the parameter called lambda. You can manually set this value or select it through cross-validation.
We will now look at an example of the use of smoothing splines with the “Clothing” dataset from the “Ecdat” package. We want to predict “tsales” based on the use of innovation in the stores. Below is some initial code.
library(Ecdat)data(Clothing)
str(Clothing)## 'data.frame': 400 obs. of 13 variables:
## $ tsales : int 750000 1926395 1250000 694227 750000 400000 1300000 495340 1200000 495340 ...
## $ sales : num 4412 4281 4167 2670 15000 ...
## $ margin : num 41 39 40 40 44 41 39 28 41 37 ...
## $ nown : num 1 2 1 1 2 ...
## $ nfull : num 1 2 2 1 1.96 ...
## $ npart : num 1 3 2.22 1.28 1.28 ...
## $ naux : num 1.54 1.54 1.41 1.37 1.37 ...
## $ hoursw : int 76 192 114 100 104 72 161 80 158 87 ...
## $ hourspw: num 16.8 22.5 17.2 21.5 15.7 ...
## $ inv1 : num 17167 17167 292857 22207 22207 ...
## $ inv2 : num 27177 27177 71571 15000 10000 ...
## $ ssize : int 170 450 300 260 50 90 400 100 450 75 ...
## $ start : num 41 39 40 40 44 41 39 28 41 37 ...We are going to create three models. Model one will have 70 degrees of freedom, model two will have 7, and model three will have the number of degrees of freedom are determined through cross-validation. Below is the code.
fit1<-smooth.spline(Clothing$inv2,Clothing$tsales,df=57)
fit2<-smooth.spline(Clothing$inv2,Clothing$tsales,df=7)
fit3<-smooth.spline(Clothing$inv2,Clothing$tsales,cv=T)## Warning in smooth.spline(Clothing$inv2, Clothing$tsales, cv = T): cross-
## validation with non-unique 'x' values seems doubtful(data.frame(fit1$df,fit2$df,fit3$df))## fit1.df fit2.df fit3.df
## 1 57 7.000957 2.791762In the code above we used the “smooth.spline” function which comes with base r.Notice that we did not use the same coding syntax as the “lm” function calls for. The code above also indicates the degrees of freedom for each model. You can see that for “fit3” the cross-validation determine that 2.79 was the most appropriate degrees of freedom. In addition, if you type in the following code.
sapply(data.frame(fit1$x,fit2$x,fit3$x),length)## fit1.x fit2.x fit3.x
## 73 73 73You will see that there are only 73 data points in each model. The “Clothing” dataset has 400 examples in it. The reason for this reduction is that the “smooth.spline” function only takes unique values from the original dataset. As such, though there are 400 examples in the dataset only 73 of them are unique.
Next, we plot our data and add regression lines
plot(Clothing$inv2,Clothing$tsales)
lines(fit1,col='red',lwd=3)
lines(fit2,col='green',lwd=3)
lines(fit3,col='blue',lwd=3)
legend('topright',lty=1,col=c('red','green','blue'),c("df = 57",'df=7','df=CV 2.8'))
You can see that as the degrees of freedom increase so does the flexibility in the line. The advantage of smoothing splines is to have a more flexible way to assess the characteristics of a dataset.
Polynomial Spline Regression in R
Normally, when least squares regression is used, you fit one line to the model. However, sometimes you may want enough flexibility that you fit different lines over different regions of your independent variable. This process of fitting different lines over different regions of X is known as Regression Splines.
How this works is that there are different coefficient values based on the regions of X. As the researcher, you can set the cutoff points for each region. The cutoff point is called a “knot.” The more knots you use the more flexible the model becomes because there are fewer data points with each range allowing for more variability.
We will now go through an example of polynomial regression splines. Remeber that polynomial means that we will have a curved line as we are using higher order polynomials. Our goal will be to predict total sales based on the amount of innovation a store employs. We will use the “Ecdat” package and the “Clothing” dataset. In addition, we will need the “splines” package. The code is as follows.
library(splines);library(Ecdat)data(Clothing)We will now fit our model. We must indicate the number and placement of the knots. This is commonly down at the 25th 50th and 75th percentile. Below is the code
fit<-lm(tsales~bs(inv2,knots = c(12000,60000,150000)),data = Clothing)In the code above we used the traditional “lm” function to set the model. However, we also used the “bs” function which allows us to create our spline regression model. The argument “knots” was set to have three different values. Lastly, the dataset was indicated.
Remember that the default spline model in R is a third-degree polynomial. This is because it is hard for the eye to detect the discontinuity at the knots.
We now need X values that we can use for prediction purposes. In the code below we first find the range of the “inv2” variable. We then create a grid that includes all the possible values of “inv2” in increments of 1. Lastly, we use the “predict” function to develop the prediction model. We set the “se” argument to true as we will need this information. The code is below.
inv2lims<-range(Clothing$inv2)
inv2.grid<-seq(from=inv2lims[1],to=inv2lims[2])
pred<-predict(fit,newdata=list(inv2=inv2.grid),se=T)We are now ready to plot our model. The code below graphs the model and includes the regression line (red), confidence interval (green), as well as the location of each knot (blue)
plot(Clothing$inv2,Clothing$tsales,main="Regression Spline Plot")
lines(inv2.grid,pred$fit,col='red',lwd=3)
lines(inv2.grid,pred$fit+2*pred$se.fit,lty="dashed",lwd=2,col='green')
lines(inv2.grid,pred$fit-2*pred$se.fit,lty="dashed",lwd=2,col='green')
segments(12000,0,x1=12000,y1=5000000,col='blue' )
segments(60000,0,x1=60000,y1=5000000,col='blue' )
segments(150000,0,x1=150000,y1=5000000,col='blue' )
When this model was created it was essentially three models connected. Model on goes from the first blue line to the second. Model 2 goes form the second blue line to the third and model three was from the third blue line until the end. This kind of flexibility is valuable in understanding nonlinear relationship
Logistic Polynomial Regression in R
Polynomial regression is used when you want to develop a regression model that is not linear. It is common to use this method when performing traditional least squares regression. However, it is also possible to use polynomial regression when the dependent variable is categorical. As such, in this post, we will go through an example of logistic polynomial regression.
Specifically, we will use the “Clothing” dataset from the “Ecdat” package. We will divide the “tsales” dependent variable into two categories to run the analysis. Below is the code to get started.
library(Ecdat)data(Clothing)There is little preparation for this example. Below is the code for the model
fitglm<-glm(I(tsales>900000)~poly(inv2,4),data=Clothing,family = binomial)Here is what we did
1. We created an object called “fitglm” to save our results
2. We used the “glm” function to process the model
3. We used the “I” function. This told R to process the information inside the parentheses as is. As such, we did not have to make a new variable in which we split the “tsales” variable. Simply, if sales were greater than 900000 it was code 1 and 0 if less than this amount.
4. Next, we set the information for the independent variable. We used the “poly” function. Inside this function, we placed the “inv2” variable and the highest order polynomial we want to explore.
5. We set the data to “Clothing”
6. Lastly, we set the “family” argument to “binomial” which is needed for logistic regression
Below is the results
summary(fitglm)##
## Call:
## glm(formula = I(tsales > 9e+05) ~ poly(inv2, 4), family = binomial,
## data = Clothing)
##
## Deviance Residuals:
## Min 1Q Median 3Q Max
## -1.5025 -0.8778 -0.8458 1.4534 1.5681
##
## Coefficients:
## Estimate Std. Error z value Pr(>|z|)
## (Intercept) 3.074 2.685 1.145 0.2523
## poly(inv2, 4)1 641.710 459.327 1.397 0.1624
## poly(inv2, 4)2 585.975 421.723 1.389 0.1647
## poly(inv2, 4)3 259.700 178.081 1.458 0.1448
## poly(inv2, 4)4 73.425 44.206 1.661 0.0967 .
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
##
## (Dispersion parameter for binomial family taken to be 1)
##
## Null deviance: 521.57 on 399 degrees of freedom
## Residual deviance: 493.51 on 395 degrees of freedom
## AIC: 503.51
##
## Number of Fisher Scoring iterations: 13It appears that only the 4th-degree polynomial is significant and barely at that. We will now find the range of our independent variable “inv2” and make a grid from this information. Doing this will allow us to run our model using the full range of possible values for our independent variable.
inv2lims<-range(Clothing$inv2)
inv2.grid<-seq(from=inv2lims[1],to=inv2lims[2])The “inv2lims” object has two values. The lowest value in “inv2” and the highest value. These values serve as the highest and lowest values in our “inv2.grid” object. This means that we have values started at 350 and going to 400000 by 1 in a grid to be used as values for “inv2” in our prediction model. Below is our prediction model.
predsglm<-predict(fitglm,newdata=list(inv2=inv2.grid),se=T,type="response")Next, we need to calculate the probabilities that a given value of “inv2” predicts a store has “tsales” greater than 900000. The equation is as follows.
pfit<-exp(predsglm$fit)/(1+exp(predsglm$fit))Graphing this leads to interesting insights. Below is the code
plot(pfit)
You can see the curves in the line from the polynomial expression. As it appears. As inv2 increase the probability increase until the values fall between 125000 and 200000. This is interesting, to say the least.
We now need to plot the actual model. First, we need to calculate the confidence intervals. This is done with the code below.
se.bandsglm.logit<-cbind(predsglm$fit+2*predsglm$se.fit,predsglm$fit-2*predsglm$se.fit)
se.bandsglm<-exp(se.bandsglm.logit)/(1+exp(se.bandsglm.logit))The ’se.bandsglm” object contains the log odds of each example and the “se.bandsglm” has the probabilities. Now we plot the results
plot(Clothing$inv2,I(Clothing$tsales>900000),xlim=inv2lims,type='n')
points(jitter(Clothing$inv2),I((Clothing$tsales>900000)),cex=2,pch='|',col='darkgrey')
lines(inv2.grid,pfit,lwd=4)
matlines(inv2.grid,se.bandsglm,col="green",lty=6,lwd=6) In the code above we did the following.
In the code above we did the following.
1. We plotted our dependent and independent variables. However, we set the argument “type” to n which means nothing. This was done so we can add the information step-by-step.
2. We added the points. This was done using the “points” function. The “jitter” function just helps to spread the information out. The other arguments (cex, pch, col) our for aesthetics and our optional.
3. We add our logistic polynomial line based on our independent variable grid and the “pfit” object which has all of the predicted probabilities.
4. Last, we add the confidence intervals using the “matlines” function. Which includes the grid object as well as the “se.bandsglm” information.
You can see that these results are similar to when we only graphed the “pfit” information. However, we also add the confidence intervals. You can see the same dip around 125000-200000 were there is also a larger confidence interval. if you look at the plot you can see that there are fewer data points in this range which may be what is making the intervals wider.
Conclusion
Logistic polynomial regression allows the regression line to have more curves to it if it is necessary. This is useful for fitting data that is non-linear in nature.
Polynomial Regression in R
Polynomial regression is one of the easiest ways to fit a non-linear line to a data set. This is done through the use of higher order polynomials such as cubic, quadratic, etc to one or more predictor variables in a model.
Generally, polynomial regression is used for one predictor and one outcome variable. When there are several predictor variables it is more common to use generalized additive modeling/ In this post, we will use the “Clothing” dataset from the “Ecdat” package to predict total sales with the use of polynomial regression. Below is some initial code.
library(Ecdat)data(Clothing) str(Clothing)
## 'data.frame': 400 obs. of 13 variables:
## $ tsales : int 750000 1926395 1250000 694227 750000 400000 1300000 495340 1200000 495340 ...
## $ sales : num 4412 4281 4167 2670 15000 ...
## $ margin : num 41 39 40 40 44 41 39 28 41 37 ...
## $ nown : num 1 2 1 1 2 ...
## $ nfull : num 1 2 2 1 1.96 ...
## $ npart : num 1 3 2.22 1.28 1.28 ...
## $ naux : num 1.54 1.54 1.41 1.37 1.37 ...
## $ hoursw : int 76 192 114 100 104 72 161 80 158 87 ...
## $ hourspw: num 16.8 22.5 17.2 21.5 15.7 ...
## $ inv1 : num 17167 17167 292857 22207 22207 ...
## $ inv2 : num 27177 27177 71571 15000 10000 ...
## $ ssize : int 170 450 300 260 50 90 400 100 450 75 ...
## $ start : num 41 39 40 40 44 41 39 28 41 37 ...We are going to use the “inv2” variable as our predictor. This variable measures the investment in automation by a particular store. We will now run our polynomial regression model.
fit<-lm(tsales~poly(inv2,5),data = Clothing)
summary(fit)##
## Call:
## lm(formula = tsales ~ poly(inv2, 5), data = Clothing)
##
## Residuals:
## Min 1Q Median 3Q Max
## -946668 -336447 -96763 184927 3599267
##
## Coefficients:
## Estimate Std. Error t value Pr(>|t|)
## (Intercept) 833584 28489 29.259 < 2e-16 ***
## poly(inv2, 5)1 2391309 569789 4.197 3.35e-05 ***
## poly(inv2, 5)2 -665063 569789 -1.167 0.2438
## poly(inv2, 5)3 49793 569789 0.087 0.9304
## poly(inv2, 5)4 1279190 569789 2.245 0.0253 *
## poly(inv2, 5)5 -341189 569789 -0.599 0.5497
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
##
## Residual standard error: 569800 on 394 degrees of freedom
## Multiple R-squared: 0.05828, Adjusted R-squared: 0.04633
## F-statistic: 4.876 on 5 and 394 DF, p-value: 0.0002428The code above should be mostly familiar. We use the “lm” function as normal for regression. However, we then used the “poly” function on the “inv2” variable. What this does is runs our model 5 times (5 is the number next to “inv2”). Each time a different polynomial is used from 1 (no polynomial) to 5 (5th order polynomial). The results indicate that the 4th-degree polynomial is significant.
We now will prepare a visual of the results but first, there are several things we need to prepare. First, we want to find what the range of our predictor variable “inv2” is and we will save this information in a grade. The code is below.
inv2lims<-range(Clothing$inv2)Second, we need to create a grid that has all the possible values of “inv2” from the lowest to the highest broken up in intervals of one. Below is the code.
inv2.grid<-seq(from=inv2lims[1],to=inv2lims[2])We now have a dataset with almost 400000 data points in the “inv2.grid” object through this approach. We will now use these values to predict “tsales.” We also want the standard errors so we se “se” to TRUE
preds<-predict(fit,newdata=list(inv2=inv2.grid),se=TRUE)We now need to find the confidence interval for our regression line. This is done by making a dataframe that takes the predicted fit adds or subtracts 2 and multiples this number by the standard error as shown below.
se.bands<-cbind(preds$fit+2*preds$se.fit,preds$fit-2*preds$se.fit)With these steps completed, we are ready to create our civilization.
To make our visual, we use the “plot” function on the predictor and outcome. Doing this gives us a plot without a regression line. We then use the “lines” function to add the polynomial regression line, however, this line is based on the “inv2.grid” object (40,000 observations) and our predictions. Lastly, we use the “matlines” function to add the confidence intervals we found and stored in the “se.bands” object.
plot(Clothing$inv2,Clothing$tsales)
lines(inv2.grid,preds$fit,lwd=4,col='blue')
matlines(inv2.grid,se.bands,lwd = 4,col = "yellow",lty=4)
Conclusion
You can clearly see the curvature of the line. Which helped to improve model fit. Now any of you can tell that we are fitting this line to mostly outliers. This is one reason we the standard error gets wider and wider it is because there are fewer and fewer observations on which to base it. However, for demonstration purposes, this is a clear example of the power of polynomial regression.
Partial Least Squares Regression in R
Partial least squares regression is a form of regression that involves the development of components of the original variables in a supervised way. What this means is that the dependent variable is used to help create the new components form the original variables. This means that when pls is used the linear combination of the new features helps to explain both the independent and dependent variables in the model.
In this post, we will use predict “income” in the “Mroz” dataset using pls. Below is some initial code.
library(pls);library(Ecdat)data("Mroz")
str(Mroz)## 'data.frame': 753 obs. of 18 variables:
## $ work : Factor w/ 2 levels "yes","no": 2 2 2 2 2 2 2 2 2 2 ...
## $ hoursw : int 1610 1656 1980 456 1568 2032 1440 1020 1458 1600 ...
## $ child6 : int 1 0 1 0 1 0 0 0 0 0 ...
## $ child618 : int 0 2 3 3 2 0 2 0 2 2 ...
## $ agew : int 32 30 35 34 31 54 37 54 48 39 ...
## $ educw : int 12 12 12 12 14 12 16 12 12 12 ...
## $ hearnw : num 3.35 1.39 4.55 1.1 4.59 ...
## $ wagew : num 2.65 2.65 4.04 3.25 3.6 4.7 5.95 9.98 0 4.15 ...
## $ hoursh : int 2708 2310 3072 1920 2000 1040 2670 4120 1995 2100 ...
## $ ageh : int 34 30 40 53 32 57 37 53 52 43 ...
## $ educh : int 12 9 12 10 12 11 12 8 4 12 ...
## $ wageh : num 4.03 8.44 3.58 3.54 10 ...
## $ income : int 16310 21800 21040 7300 27300 19495 21152 18900 20405 20425 ...
## $ educwm : int 12 7 12 7 12 14 14 3 7 7 ...
## $ educwf : int 7 7 7 7 14 7 7 3 7 7 ...
## $ unemprate : num 5 11 5 5 9.5 7.5 5 5 3 5 ...
## $ city : Factor w/ 2 levels "no","yes": 1 2 1 1 2 2 1 1 1 1 ...
## $ experience: int 14 5 15 6 7 33 11 35 24 21 ...First, we must prepare our data by dividing it into a training and test set. We will do this by doing a 50/50 split of the data.
set.seed(777)
train<-sample(c(T,F),nrow(Mroz),rep=T) #50/50 train/test split
test<-(!train)In the code above we set the “set.seed function in order to assure reduplication. Then we created the “train” object and used the “sample” function to make a vector with ‘T’ and ‘F’ based on the number of rows in “Mroz”. Lastly, we created the “test”” object base don everything that is not in the “train” object as that is what the exclamation point is for.
Now we create our model using the “plsr” function from the “pls” package and we will examine the results using the “summary” function. We will also scale the data since this the scale affects the development of the components and use cross-validation. Below is the code.
set.seed(777)
pls.fit<-plsr(income~.,data=Mroz,subset=train,scale=T,validation="CV")
summary(pls.fit)## Data: X dimension: 392 17
## Y dimension: 392 1
## Fit method: kernelpls
## Number of components considered: 17
##
## VALIDATION: RMSEP
## Cross-validated using 10 random segments.
## (Intercept) 1 comps 2 comps 3 comps 4 comps 5 comps 6 comps
## CV 11218 8121 6701 6127 5952 5886 5857
## adjCV 11218 8114 6683 6108 5941 5872 5842
## 7 comps 8 comps 9 comps 10 comps 11 comps 12 comps 13 comps
## CV 5853 5849 5854 5853 5853 5852 5852
## adjCV 5837 5833 5837 5836 5836 5835 5835
## 14 comps 15 comps 16 comps 17 comps
## CV 5852 5852 5852 5852
## adjCV 5835 5835 5835 5835
##
## TRAINING: % variance explained
## 1 comps 2 comps 3 comps 4 comps 5 comps 6 comps 7 comps
## X 17.04 26.64 37.18 49.16 59.63 64.63 69.13
## income 49.26 66.63 72.75 74.16 74.87 75.25 75.44
## 8 comps 9 comps 10 comps 11 comps 12 comps 13 comps 14 comps
## X 72.82 76.06 78.59 81.79 85.52 89.55 92.14
## income 75.49 75.51 75.51 75.52 75.52 75.52 75.52
## 15 comps 16 comps 17 comps
## X 94.88 97.62 100.00
## income 75.52 75.52 75.52The printout includes the root mean squared error for each of the components in the VALIDATION section as well as the variance explained in the TRAINING section. There are 17 components because there are 17 independent variables. You can see that after component 3 or 4 there is little improvement in the variance explained in the dependent variable. Below is the code for the plot of these results. It requires the use of the “validationplot” function with the “val.type” argument set to “MSEP” Below is the code
validationplot(pls.fit,val.type = "MSEP")
We will do the predictions with our model. We use the “predict” function, use our “Mroz” dataset but only those index in the “test” vector and set the components to three based on our previous plot. Below is the code.
set.seed(777)
pls.pred<-predict(pls.fit,Mroz[test,],ncomp=3)After this, we will calculate the mean squared error. This is done by subtracting the results of our predicted model from the dependent variable of the test set. We then square this information and calculate the mean. Below is the code
mean((pls.pred-Mroz$income[test])^2)## [1] 63386682As you know, this information is only useful when compared to something else. Therefore, we will run the data with a tradition least squares regression model and compare the results.
set.seed(777)
lm.fit<-lm(income~.,data=Mroz,subset=train)
lm.pred<-predict(lm.fit,Mroz[test,])
mean((lm.pred-Mroz$income[test])^2)## [1] 59432814The least squares model is slightly better then our partial least squares model but if we look at the model we see several variables that are not significant. We will remove these see what the results are
summary(lm.fit)##
## Call:
## lm(formula = income ~ ., data = Mroz, subset = train)
##
## Residuals:
## Min 1Q Median 3Q Max
## -20131 -2923 -1065 1670 36246
##
## Coefficients:
## Estimate Std. Error t value Pr(>|t|)
## (Intercept) -1.946e+04 3.224e+03 -6.036 3.81e-09 ***
## workno -4.823e+03 1.037e+03 -4.651 4.59e-06 ***
## hoursw 4.255e+00 5.517e-01 7.712 1.14e-13 ***
## child6 -6.313e+02 6.694e+02 -0.943 0.346258
## child618 4.847e+02 2.362e+02 2.052 0.040841 *
## agew 2.782e+02 8.124e+01 3.424 0.000686 ***
## educw 1.268e+02 1.889e+02 0.671 0.502513
## hearnw 6.401e+02 1.420e+02 4.507 8.79e-06 ***
## wagew 1.945e+02 1.818e+02 1.070 0.285187
## hoursh 6.030e+00 5.342e-01 11.288 < 2e-16 ***
## ageh -9.433e+01 7.720e+01 -1.222 0.222488
## educh 1.784e+02 1.369e+02 1.303 0.193437
## wageh 2.202e+03 8.714e+01 25.264 < 2e-16 ***
## educwm -4.394e+01 1.128e+02 -0.390 0.697024
## educwf 1.392e+02 1.053e+02 1.322 0.186873
## unemprate -1.657e+02 9.780e+01 -1.694 0.091055 .
## cityyes -3.475e+02 6.686e+02 -0.520 0.603496
## experience -1.229e+02 4.490e+01 -2.737 0.006488 **
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
##
## Residual standard error: 5668 on 374 degrees of freedom
## Multiple R-squared: 0.7552, Adjusted R-squared: 0.744
## F-statistic: 67.85 on 17 and 374 DF, p-value: < 2.2e-16set.seed(777)
lm.fit<-lm(income~work+hoursw+child618+agew+hearnw+hoursh+wageh+experience,data=Mroz,subset=train)
lm.pred<-predict(lm.fit,Mroz[test,])
mean((lm.pred-Mroz$income[test])^2)## [1] 57839715As you can see the error decreased even more which indicates that the least squares regression model is superior to the partial least squares model. In addition, the partial least squares model is much more difficult to explain because of the use of components. As such, the least squares model is the favored one.
Principal Component Regression in R
This post will explain and provide an example of principal component regression (PCR). Principal component regression involves having the model construct components from the independent variables that are a linear combination of the independent variables. This is similar to principal component analysis but the components are designed in a way to best explain the dependent variable. Doing this often allows you to use fewer variables in your model and usually improves the fit of your model as well.
Since PCR is based on principal component analysis it is an unsupervised method, which means the dependent variable has no influence on the development of the components. As such, there are times when the components that are developed may not be beneficial for explaining the dependent variable.
Our example will use the “Mroz” dataset from the “Ecdat” package. Our goal will be to predict “income” based on the variables in the dataset. Below is the initial code
library(pls);library(Ecdat)data(Mroz)
str(Mroz)## 'data.frame': 753 obs. of 18 variables:
## $ work : Factor w/ 2 levels "yes","no": 2 2 2 2 2 2 2 2 2 2 ...
## $ hoursw : int 1610 1656 1980 456 1568 2032 1440 1020 1458 1600 ...
## $ child6 : int 1 0 1 0 1 0 0 0 0 0 ...
## $ child618 : int 0 2 3 3 2 0 2 0 2 2 ...
## $ agew : int 32 30 35 34 31 54 37 54 48 39 ...
## $ educw : int 12 12 12 12 14 12 16 12 12 12 ...
## $ hearnw : num 3.35 1.39 4.55 1.1 4.59 ...
## $ wagew : num 2.65 2.65 4.04 3.25 3.6 4.7 5.95 9.98 0 4.15 ...
## $ hoursh : int 2708 2310 3072 1920 2000 1040 2670 4120 1995 2100 ...
## $ ageh : int 34 30 40 53 32 57 37 53 52 43 ...
## $ educh : int 12 9 12 10 12 11 12 8 4 12 ...
## $ wageh : num 4.03 8.44 3.58 3.54 10 ...
## $ income : int 16310 21800 21040 7300 27300 19495 21152 18900 20405 20425 ...
## $ educwm : int 12 7 12 7 12 14 14 3 7 7 ...
## $ educwf : int 7 7 7 7 14 7 7 3 7 7 ...
## $ unemprate : num 5 11 5 5 9.5 7.5 5 5 3 5 ...
## $ city : Factor w/ 2 levels "no","yes": 1 2 1 1 2 2 1 1 1 1 ...
## $ experience: int 14 5 15 6 7 33 11 35 24 21 ...Our first step is to divide our dataset into a train and test set. We will do a simple 50/50 split for this demonstration.
train<-sample(c(T,F),nrow(Mroz),rep=T) #50/50 train/test split
test<-(!train)In the code above we use the “sample” function to create a “train” index based on the number of rows in the “Mroz” dataset. Basically, R is making a vector that randomly assigns different rows in the “Mroz” dataset to be marked as True or False. Next, we use the “train” vector and we assign everything or every number that is not in the “train” vector to the test vector by using the exclamation mark.
We are now ready to develop our model. Below is the code
set.seed(777)
pcr.fit<-pcr(income~.,data=Mroz,subset=train,scale=T,validation="CV")To make our model we use the “pcr” function from the “pls” package. The “subset” argument tells r to use the “train” vector to select examples from the “Mroz” dataset. The “scale” argument makes sure everything is measured the same way. This is important when using a component analysis tool as variables with different scale have a different influence on the components. Lastly, the “validation” argument enables cross-validation. This will help us to determine the number of components to use for prediction. Below is the results of the model using the “summary” function.
summary(pcr.fit)## Data: X dimension: 381 17
## Y dimension: 381 1
## Fit method: svdpc
## Number of components considered: 17
##
## VALIDATION: RMSEP
## Cross-validated using 10 random segments.
## (Intercept) 1 comps 2 comps 3 comps 4 comps 5 comps 6 comps
## CV 12102 11533 11017 9863 9884 9524 9563
## adjCV 12102 11534 11011 9855 9878 9502 9596
## 7 comps 8 comps 9 comps 10 comps 11 comps 12 comps 13 comps
## CV 9149 9133 8811 8527 7265 7234 7120
## adjCV 9126 9123 8798 8877 7199 7172 7100
## 14 comps 15 comps 16 comps 17 comps
## CV 7118 7141 6972 6992
## adjCV 7100 7123 6951 6969
##
## TRAINING: % variance explained
## 1 comps 2 comps 3 comps 4 comps 5 comps 6 comps 7 comps
## X 21.359 38.71 51.99 59.67 65.66 71.20 76.28
## income 9.927 19.50 35.41 35.63 41.28 41.28 46.75
## 8 comps 9 comps 10 comps 11 comps 12 comps 13 comps 14 comps
## X 80.70 84.39 87.32 90.15 92.65 95.02 96.95
## income 47.08 50.98 51.73 68.17 68.29 68.31 68.34
## 15 comps 16 comps 17 comps
## X 98.47 99.38 100.00
## income 68.48 70.29 70.39There is a lot of information here.The VALIDATION: RMSEP section gives you the root mean squared error of the model broken down by component. The TRAINING section is similar the printout of any PCA but it shows the amount of cumulative variance of the components, as well as the variance, explained for the dependent variable “income.” In this model, we are able to explain up to 70% of the variance if we use all 17 components.
We can graph the MSE using the “validationplot” function with the argument “val.type” set to “MSEP”. The code is below.
validationplot(pcr.fit,val.type = "MSEP")
How many components to pick is subjective, however, there is almost no improvement beyond 13 so we will use 13 components in our prediction model and we will calculate the means squared error.
set.seed(777)
pcr.pred<-predict(pcr.fit,Mroz[test,],ncomp=13)
mean((pcr.pred-Mroz$income[test])^2)## [1] 48958982MSE is what you would use to compare this model to other models that you developed. Below is the performance of a least squares regression model
set.seed(777)
lm.fit<-lm(income~.,data=Mroz,subset=train)
lm.pred<-predict(lm.fit,Mroz[test,])
mean((lm.pred-Mroz$income[test])^2)## [1] 47794472If you compare the MSE the least squares model performs slightly better than the PCR one. However, there are a lot of non-significant features in the model as shown below.
summary(lm.fit)##
## Call:
## lm(formula = income ~ ., data = Mroz, subset = train)
##
## Residuals:
## Min 1Q Median 3Q Max
## -27646 -3337 -1387 1860 48371
##
## Coefficients:
## Estimate Std. Error t value Pr(>|t|)
## (Intercept) -2.215e+04 3.987e+03 -5.556 5.35e-08 ***
## workno -3.828e+03 1.316e+03 -2.909 0.00385 **
## hoursw 3.955e+00 7.085e-01 5.582 4.65e-08 ***
## child6 5.370e+02 8.241e+02 0.652 0.51512
## child618 4.250e+02 2.850e+02 1.491 0.13673
## agew 1.962e+02 9.849e+01 1.992 0.04709 *
## educw 1.097e+02 2.276e+02 0.482 0.63013
## hearnw 9.835e+02 2.303e+02 4.270 2.50e-05 ***
## wagew 2.292e+02 2.423e+02 0.946 0.34484
## hoursh 6.386e+00 6.144e-01 10.394 < 2e-16 ***
## ageh -1.284e+01 9.762e+01 -0.132 0.89542
## educh 1.460e+02 1.592e+02 0.917 0.35982
## wageh 2.083e+03 9.930e+01 20.978 < 2e-16 ***
## educwm 1.354e+02 1.335e+02 1.014 0.31115
## educwf 1.653e+02 1.257e+02 1.315 0.18920
## unemprate -1.213e+02 1.148e+02 -1.057 0.29140
## cityyes -2.064e+02 7.905e+02 -0.261 0.79421
## experience -1.165e+02 5.393e+01 -2.159 0.03147 *
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
##
## Residual standard error: 6729 on 363 degrees of freedom
## Multiple R-squared: 0.7039, Adjusted R-squared: 0.69
## F-statistic: 50.76 on 17 and 363 DF, p-value: < 2.2e-16Removing these and the MSE is almost the same for the PCR and least square models
set.seed(777)
lm.fit2<-lm(income~work+hoursw+hearnw+hoursh+wageh,data=Mroz,subset=train)
lm.pred2<-predict(lm.fit2,Mroz[test,])
mean((lm.pred2-Mroz$income[test])^2)## [1] 47968996Conclusion
Since the least squares model is simpler it is probably the superior model. PCR is strongest when there are a lot of variables involve and if there are issues with multicollinearity.
Example of Best Subset Regression in R
This post will provide an example of best subset regression. This is a topic that has been covered before in this blog. However, in the current post, we will approach this using a slightly different coding and a different dataset. We will be using the “HI” dataset from the “Ecdat” package. Our goal will be to predict the number of hours a women works based on the other variables in the dataset. Below is some initial code.
library(leaps);library(Ecdat)data(HI)
str(HI)## 'data.frame': 22272 obs. of 13 variables:
## $ whrswk : int 0 50 40 40 0 40 40 25 45 30 ...
## $ hhi : Factor w/ 2 levels "no","yes": 1 1 2 1 2 2 2 1 1 1 ...
## $ whi : Factor w/ 2 levels "no","yes": 1 2 1 2 1 2 1 1 2 1 ...
## $ hhi2 : Factor w/ 2 levels "no","yes": 1 1 2 2 2 2 2 1 1 2 ...
## $ education : Ord.factor w/ 6 levels "<9years"<"9-11years"<..: 4 4 3 4 2 3 5 3 5 4 ...
## $ race : Factor w/ 3 levels "white","black",..: 1 1 1 1 1 1 1 1 1 1 ...
## $ hispanic : Factor w/ 2 levels "no","yes": 1 1 1 1 1 1 1 1 1 1 ...
## $ experience: num 13 24 43 17 44.5 32 14 1 4 7 ...
## $ kidslt6 : int 2 0 0 0 0 0 0 1 0 1 ...
## $ kids618 : int 1 1 0 1 0 0 0 0 0 0 ...
## $ husby : num 12 1.2 31.3 9 0 ...
## $ region : Factor w/ 4 levels "other","northcentral",..: 2 2 2 2 2 2 2 2 2 2 ...
## $ wght : int 214986 210119 219955 210317 219955 208148 213615 181960 214874 214874 ...To develop a model we use the “regsubset” function from the “leap” package. Most of the coding is the same as linear regression. The only difference is the “nvmax” argument which is set to 13. The default setting for “nvmax” is 8. This is good if you only have 8 variables. However, the results from the “str” function indicate that we have 13 functions. Therefore, we need to set the “nvmax” argument to 13 instead of the default value of 8 in order to be sure to include all variables. Below is the code
regfit.full<-regsubsets(whrswk~.,HI, nvmax = 13)We can look at the results with the “summary” function. For space reasons, the code is shown but the results will not be shown here.
summary(regfit.full)If you run the code above in your computer you will 13 columns that are named after the variables created. A star in a column means that that variable is included in the model. To the left is the numbers 1-13 which. One means one variable in the model two means two variables in the model etc.
Our next step is to determine which of these models is the best. First, we need to decide what our criteria for inclusion will be. Below is a list of available fit indices.
names(summary(regfit.full))## [1] "which" "rsq" "rss" "adjr2" "cp" "bic" "outmat" "obj"For our purposes, we will use “rsq” (r-square) and “bic” “Bayesian Information Criteria.” In the code below we are going to save the values for these two fit indices in their own objects.
rsq<-summary(regfit.full)$rsq
bic<-summary(regfit.full)$bicNow let’s plot them
plot(rsq,type='l',main="R-Square",xlab="Number of Variables")
plot(bic,type='l',main="BIC",xlab="Number of Variables")
You can see that for r-square the values increase and for BIC the values decrease. We will now make both of these plots again but we will have r tell the optimal number of variables when considering each model index. For we use the “which” function to determine the max r-square and the minimum BIC
which.max(rsq)## [1] 13which.min(bic)## [1] 12The model with the best r-square is the one with 13 variables. This makes sense as r-square always improves as you add variables. Since this is a demonstration we will not correct for this. For BIC the lowest values was for 12 variables. We will now plot this information and highlight the best model in the plot using the “points” function, which allows you to emphasis one point in a graph
plot(rsq,type='l',main="R-Square with Best Model Highlighted",xlab="Number of Variables")
points(13,(rsq[13]),col="blue",cex=7,pch=20)
plot(bic,type='l',main="BIC with Best Model Highlighted",xlab="Number of Variables")
points(12,(bic[12]),col="blue",cex=7,pch=20)
Since BIC calls for only 12 variables it is simpler than the r-square recommendation of 13. Therefore, we will fit our final model using the BIC recommendation of 12. Below is the code.
coef(regfit.full,12)## (Intercept) hhiyes whiyes
## 30.31321796 1.16940604 18.25380263
## education.L education^4 education^5
## 6.63847641 1.54324869 -0.77783663
## raceblack hispanicyes experience
## 3.06580207 -1.33731802 -0.41883100
## kidslt6 kids618 husby
## -6.02251640 -0.82955827 -0.02129349
## regionnorthcentral
## 0.94042820So here is our final model. This is what we would use for our test set.
Conclusion
Best subset regression provides the researcher with insights into every possible model as well as clues as to which model is at least statistically superior. This knowledge can be used for developing models for data science applications.
High Dimensionality Regression
There are times when least squares regression is not able to provide accurate predictions or explanation in an object. One example in which least scares regression struggles with a small sample size. By small, we mean when the total number of variables is greater than the sample size. Another term for this is high dimensions which means more variables than examples in the dataset
This post will explain the consequences of what happens when high dimensions is a problem and also how to address the problem.
Inaccurate measurements
One problem with high dimensions in regression is that the results for the various metrics are overfitted to the data. Below is an example using the “attitude” dataset. There are 2 variables and 3 examples for developing a model. This is not strictly high dimensions but it is an example of a small sample size.
data("attitude")
reg1 <- lm(complaints[1:3]~rating[1:3],data=attitude[1:3])
summary(reg1)##
## Call:
## lm(formula = complaints[1:3] ~ rating[1:3], data = attitude[1:3])
##
## Residuals:
## 1 2 3
## 0.1026 -0.3590 0.2564
##
## Coefficients:
## Estimate Std. Error t value Pr(>|t|)
## (Intercept) 21.95513 1.33598 16.43 0.0387 *
## rating[1:3] 0.67308 0.02221 30.31 0.0210 *
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
##
## Residual standard error: 0.4529 on 1 degrees of freedom
## Multiple R-squared: 0.9989, Adjusted R-squared: 0.9978
## F-statistic: 918.7 on 1 and 1 DF, p-value: 0.021With only 3 data points the fit is perfect. You can also examine the mean squared error of the model. Below is a function for this followed by the results
mse <- function(sm){
mean(sm$residuals^2)}
mse(reg1)## [1] 0.06837607Almost no error. Lastly, let’s look at a visual of the model
with(attitude[1:3],plot(complaints[1:3]~ rating[1:3]))
title(main = "Sample Size 3")
abline(lm(complaints[1:3]~rating[1:3],data = attitude))
You can see that the regression line goes almost perfectly through each data point. If we tried to use this model on the test set in a real data science problem there would be a huge amount of bias. Now we will rerun the analysis this time with the full sample.
reg2<- lm(complaints~rating,data=attitude)
summary(reg2)##
## Call:
## lm(formula = complaints ~ rating, data = attitude)
##
## Residuals:
## Min 1Q Median 3Q Max
## -13.3880 -6.4553 -0.2997 6.1462 13.3603
##
## Coefficients:
## Estimate Std. Error t value Pr(>|t|)
## (Intercept) 8.2445 7.6706 1.075 0.292
## rating 0.9029 0.1167 7.737 1.99e-08 ***
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
##
## Residual standard error: 7.65 on 28 degrees of freedom
## Multiple R-squared: 0.6813, Adjusted R-squared: 0.6699
## F-statistic: 59.86 on 1 and 28 DF, p-value: 1.988e-08You can clearly see a huge reduction in the r-square from .99 to .68. Next, is the mean-square error
mse(reg2)## [1] 54.61425The error has increased a great deal. Lastly, we fit the regression line
with(attitude,plot(complaints~ rating))
title(main = "Full Sample Size")
abline(lm(complaints~rating,data = attitude))
Naturally, the second model is more likely to perform better with a test set. The problem is that least squares regression is too flexible when the number of features is greater than or equal to the number of examples in a dataset.
What to Do?
If least squares regression must be used. One solution to overcoming high dimensionality is to use some form of regularization regression such as ridge, lasso, or elastic net. Any of these regularization approaches will help to reduce the number of variables or dimensions in the final model through the use of shrinkage.
However, keep in mind that no matter what you do as the number of dimensions increases so does the r-square even if the variable is useless. This is known as the curse of dimensionality. Again, regularization can help with this.
Remember that with a large number of dimensions there are normally several equally acceptable models. To determine which is most useful depends on understanding the problem and context of the study.
Conclusion
With the ability to collect huge amounts of data has led to the growing problem of high dimensionality. One there are more features than examples it can lead to statistical errors. However, regularization is one tool for dealing with this problem.

Regression with Shrinkage Methods
One problem with least squares regression is determining what variables to keep in a model. One solution to this problem is the use of shrinkage methods. Shrinkage regression involves constraining or regularizing the coefficient estimates towards zero. The benefit of this is that it is an efficient way to either remove variables from a model or significantly reduce the influence of less important variables.
In this post, we will look at two common forms of regularization and these are.
Ridge
Ridge regression includes a tuning parameter called lambda that can be used to reduce to weak coefficients almost to zero. This shrinkage penalty helps with the bias-variance trade-off. Lambda can be set to any value from 0 to infinity. A lambda set to 0 is the same as least square regression while a lambda set to infinity will produce a null model. The technical term for lambda when ridge is used is the “l2 norm”
Finding the right value of lambda is the primary goal when using this algorithm,. Finding it involves running models with several values of lambda and seeing which returns the best results on predetermined metrics.
The primary problem with ridge regression is that it does not actually remove any variables from the model. As such, the prediction might be excellent but explanatory power is not improve if there are a large number of variables.
Lasso
Lasso regression has the same characteristics as Ridge with one exception. The one exception is the Lasso algorithm can actually remove variables by setting them to zero. This means that lasso regression models are usually superior in terms of the ability to interpret and explain them. The technical term for lambda when lasso is used is the “l1 norm.”
It is not clear when lasso or ridge is superior. Normally, if the goal is explanatory lasso is often stronger. However, if the goal is prediction, ridge may be an improvement but not always.
Conclusion
Shrinkage methods are not limited to regression. Many other forms of analysis can employ shrinkage such as artificial neural networks. Most machine learning models can accommodate shrinkage.
Generally, ridge and lasso regression is employed when you have a huge number of predictors as well as a larger dataset. The primary goal is the simplification of an overly complex model. Therefore, the shrinkage methods mentioned here are additional ways to use statistical models in regression.

Subset Selection Regression
There are many different ways in which the variables of a regression model can be selected. In this post, we look at several common ways in which to select variables or features for a regression model. In particular, we look at the following.
- Best subset regression
- Stepwise selection
Best Subset Regression
Best subset regression fits a regression model for every possible combination of variables. The “best” model can be selected based on such criteria as the adjusted r-square, BIC (Bayesian Information Criteria), etc.
The primary drawback to best subset regression is that it becomes impossible to compute the results when you have a large number of variables. Generally, when the number of variables exceeds 40 best subset regression becomes too difficult to calculate.
Stepwise Selection
Stepwise selection involves adding or taking away one variable at a time from a regression model. There are two forms of stepwise selection and they are forward and backward selection.
In forward selection, the computer starts with a null model ( a model that calculates the mean) and adds one variable at a time to the model. The variable chosen is the one the provides the best improvement to the model fit. This process reduces greatly the number of models that need to be fitted in comparison to best subset regression.
Backward selection starts the full model and removes one variable at a time based on which variable improves the model fit the most. The main problem with either forward or backward selection is that the best model may not always be selected in this process. In addition, backward selection must have a sample size that is larger than the number of variables.
Deciding Which to Choose
Best subset regression is perhaps most appropriate when you have a small number of variables to develop a model with, such as less than 40. When the number of variables grows forward or backward selection are appropriate. If the sample size is small forward selection may be a better choice. However, if the sample size is large as in the number of examples is greater than the number of variables it is now possible to use backward selection.
Conclusion
The examples here are some of the most basic ways to develop a regression model. However, these are not the only ways in which this can be done. What these examples provide is an introduction to regression model development. In addition, these models provide some sort of criteria for the addition or removal of a variable based on statistics rather than intuition.

Additive Assumption and Multiple Regression
In regression, one of the assumptions is the additive assumption. This assumption states that the influence of a predictor variable on the dependent variable is independent of any other influence. However, in practice, it is common that this assumption does not hold.
In this post, we will look at how to address violations of the additive assumption through the use of interactions in a regression model.
An interaction effect is when you have two predictor variables whose effect on the dependent variable is not the same. As such, their effect must be considered simultaneously rather than separately. This is done through the use of an interaction term. An interaction term is the product of the two predictor variables.
Let’s begin by making a regular regression model with an interaction. To do this we will use the “Carseats” data from the “ISLR” package to predict “Sales”. Below is the code.
library(ISLR);library(ggplot2)
data(Carseats)
saleslm<-lm(Sales~.,Carseats)
summary(saleslm)##
## Call:
## lm(formula = Sales ~ ., data = Carseats)
##
## Residuals:
## Min 1Q Median 3Q Max
## -2.8692 -0.6908 0.0211 0.6636 3.4115
##
## Coefficients:
## Estimate Std. Error t value Pr(>|t|)
## (Intercept) 5.6606231 0.6034487 9.380 < 2e-16 ***
## CompPrice 0.0928153 0.0041477 22.378 < 2e-16 ***
## Income 0.0158028 0.0018451 8.565 2.58e-16 ***
## Advertising 0.1230951 0.0111237 11.066 < 2e-16 ***
## Population 0.0002079 0.0003705 0.561 0.575
## Price -0.0953579 0.0026711 -35.700 < 2e-16 ***
## ShelveLocGood 4.8501827 0.1531100 31.678 < 2e-16 ***
## ShelveLocMedium 1.9567148 0.1261056 15.516 < 2e-16 ***
## Age -0.0460452 0.0031817 -14.472 < 2e-16 ***
## Education -0.0211018 0.0197205 -1.070 0.285
## UrbanYes 0.1228864 0.1129761 1.088 0.277
## USYes -0.1840928 0.1498423 -1.229 0.220
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
##
## Residual standard error: 1.019 on 388 degrees of freedom
## Multiple R-squared: 0.8734, Adjusted R-squared: 0.8698
## F-statistic: 243.4 on 11 and 388 DF, p-value: < 2.2e-16The results are rather excellent for the social sciences. The model explains 87.3% of the variance in “Sales”. The current results that we have are known as main effects. These are effects that directly influence the dependent variable. Most regression models only include main effects.
We will now examine an interaction effect between two continuous variables. Let’s see if there is an interaction between “Population” and “Income”.
saleslm1<-lm(Sales~CompPrice+Income+Advertising+Population+Price+Age+Education+US+
Urban+ShelveLoc+Population*Income, Carseats)
summary(saleslm1)##
## Call:
## lm(formula = Sales ~ CompPrice + Income + Advertising + Population +
## Price + Age + Education + US + Urban + ShelveLoc + Population *
## Income, data = Carseats)
##
## Residuals:
## Min 1Q Median 3Q Max
## -2.8699 -0.7624 0.0139 0.6763 3.4344
##
## Coefficients:
## Estimate Std. Error t value Pr(>|t|)
## (Intercept) 6.195e+00 6.436e-01 9.625 <2e-16 ***
## CompPrice 9.262e-02 4.126e-03 22.449 <2e-16 ***
## Income 7.973e-03 3.869e-03 2.061 0.0400 *
## Advertising 1.237e-01 1.107e-02 11.181 <2e-16 ***
## Population -1.811e-03 9.524e-04 -1.901 0.0580 .
## Price -9.511e-02 2.659e-03 -35.773 <2e-16 ***
## Age -4.566e-02 3.169e-03 -14.409 <2e-16 ***
## Education -2.157e-02 1.961e-02 -1.100 0.2722
## USYes -2.160e-01 1.497e-01 -1.443 0.1498
## UrbanYes 1.330e-01 1.124e-01 1.183 0.2375
## ShelveLocGood 4.859e+00 1.523e-01 31.901 <2e-16 ***
## ShelveLocMedium 1.964e+00 1.255e-01 15.654 <2e-16 ***
## Income:Population 2.879e-05 1.253e-05 2.298 0.0221 *
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
##
## Residual standard error: 1.013 on 387 degrees of freedom
## Multiple R-squared: 0.8751, Adjusted R-squared: 0.8712
## F-statistic: 226 on 12 and 387 DF, p-value: < 2.2e-16The new contribution is at the bottom of the coefficient table and is the “Income:Population” coefficient. What this means is “the increase of Sales given a one unit increase in Income and Population simultaneously” In other words the “Income:Population” coefficient looks at their combined simultaneous effect on Sales rather than just their independent effect on Sales.
This makes practical sense as well. The larger the population the more available income and vice versa. However, for our current model, the improvement in the r-squared is relatively small. The actual effect is a small increase in sales. Below is a graph of income and population by sales. Notice how the lines cross. This is a visual of what an interaction looks like. The lines are not parallel by any means.
ggplot(data=Carseats, aes(x=Income, y=Sales, group=1)) +geom_smooth(method=lm,se=F)+
geom_smooth(aes(Population,Sales), method=lm, se=F,color="black")+xlab("Income and Population")+labs(
title="Income in Blue Population in Black")We will now repeat this process but this time using a categorical variable and a continuous variable. We will look at the interaction between “US” location (categorical) and “Advertising” (continuous).
saleslm2<-lm(Sales~CompPrice+Income+Advertising+Population+Price+Age+Education+US+
Urban+ShelveLoc+US*Advertising, Carseats)
summary(saleslm2)##
## Call:
## lm(formula = Sales ~ CompPrice + Income + Advertising + Population +
## Price + Age + Education + US + Urban + ShelveLoc + US * Advertising,
## data = Carseats)
##
## Residuals:
## Min 1Q Median 3Q Max
## -2.8531 -0.7140 0.0266 0.6735 3.3773
##
## Coefficients:
## Estimate Std. Error t value Pr(>|t|)
## (Intercept) 5.6995305 0.6023074 9.463 < 2e-16 ***
## CompPrice 0.0926214 0.0041384 22.381 < 2e-16 ***
## Income 0.0159111 0.0018414 8.641 < 2e-16 ***
## Advertising 0.2130932 0.0530297 4.018 7.04e-05 ***
## Population 0.0001540 0.0003708 0.415 0.6782
## Price -0.0954623 0.0026649 -35.823 < 2e-16 ***
## Age -0.0463674 0.0031789 -14.586 < 2e-16 ***
## Education -0.0233500 0.0197122 -1.185 0.2369
## USYes -0.1057320 0.1561265 -0.677 0.4987
## UrbanYes 0.1191653 0.1127047 1.057 0.2910
## ShelveLocGood 4.8726025 0.1532599 31.793 < 2e-16 ***
## ShelveLocMedium 1.9665296 0.1259070 15.619 < 2e-16 ***
## Advertising:USYes -0.0933384 0.0537807 -1.736 0.0834 .
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
##
## Residual standard error: 1.016 on 387 degrees of freedom
## Multiple R-squared: 0.8744, Adjusted R-squared: 0.8705
## F-statistic: 224.5 on 12 and 387 DF, p-value: < 2.2e-16Again, you can see that when the store is in the US you have to also consider the advertising budget as well. When these two variables are considered there is a slight decline in sales. What this means in practice is that advertising in the US is not as beneficial as advertising outside the US.
Below you can again see a visual of the interaction effect when the lines for US yes and no cross each other in the plot below.
ggplot(data=Carseats, aes(x=Advertising, y=Sales, group = US, colour = US)) + geom_smooth(method=lm,se=F)+scale_x_continuous(limits = c(0, 25))+scale_y_continuous(limits = c(0, 25))
Lastly, we will look at an interaction effect for two categorical variables.
saleslm3<-lm(Sales~CompPrice+Income+Advertising+Population+Price+Age+Education+US+
Urban+ShelveLoc+ShelveLoc*US, Carseats)
summary(saleslm3)##
## Call:
## lm(formula = Sales ~ CompPrice + Income + Advertising + Population +
## Price + Age + Education + US + Urban + ShelveLoc + ShelveLoc *
## US, data = Carseats)
##
## Residuals:
## Min 1Q Median 3Q Max
## -2.8271 -0.6839 0.0213 0.6407 3.4537
##
## Coefficients:
## Estimate Std. Error t value Pr(>|t|)
## (Intercept) 5.8120748 0.6089695 9.544 <2e-16 ***
## CompPrice 0.0929370 0.0041283 22.512 <2e-16 ***
## Income 0.0158793 0.0018378 8.640 <2e-16 ***
## Advertising 0.1223281 0.0111143 11.006 <2e-16 ***
## Population 0.0001899 0.0003721 0.510 0.6100
## Price -0.0952439 0.0026585 -35.826 <2e-16 ***
## Age -0.0459380 0.0031830 -14.433 <2e-16 ***
## Education -0.0267021 0.0197807 -1.350 0.1778
## USYes -0.3683074 0.2379400 -1.548 0.1225
## UrbanYes 0.1438775 0.1128171 1.275 0.2030
## ShelveLocGood 4.3491643 0.2734344 15.906 <2e-16 ***
## ShelveLocMedium 1.8967193 0.2084496 9.099 <2e-16 ***
## USYes:ShelveLocGood 0.7184116 0.3320759 2.163 0.0311 *
## USYes:ShelveLocMedium 0.0907743 0.2631490 0.345 0.7303
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
##
## Residual standard error: 1.014 on 386 degrees of freedom
## Multiple R-squared: 0.8753, Adjusted R-squared: 0.8711
## F-statistic: 208.4 on 13 and 386 DF, p-value: < 2.2e-16In this case, we can see that when the store is in the US and the shelf location is good it has an effect on Sales when compared to a bad location. The plot below is a visual of this. However, it is harder to see this because the x-axis has only two categories
ggplot(data=Carseats, aes(x=US, y=Sales, group = ShelveLoc, colour = ShelveLoc)) +
geom_smooth(method=lm,se=F)Conclusion
Interactions effects are a great way to fine-tune a model, especially for explanatory purposes. Often, the change in r-square is not strong enough for prediction but can be used for nuanced understanding of the relationships among the variables.

Concepts to Consider for Model Development
When assessing how to conduct quantitative data analysis there are several factors to consider. In this post, we look at common either-or choices in data analysis. The concepts are as follows
- Parametric vs Non-Parametric
- Bias vs Variance
- Accuracy vs Interpretability
- Supervised vs Unsupervised
- Regression vs Classifications
Parametric vs Non-Parametric
Parametric models make assumptions about the shape of a function. Often, the assumption is that the function is a linear in nature such as in linear regression. Making this assumption makes it much easier to estimate the actual function of a model.
Non-parametric models do not make any assumptions about the shape of the function. This allows the function to take any shape possible. Examples of non-parametric models include decision trees, support vector machines, and artificial neural networks.
The main concern with non-parametric models is they require a huge dataset when compared to parametric models.
Bias vs Variance Tradeoff
In relation to parametric and non-parametric models is the bias-variance tradeoff. A bias model is simply a model that does not listen to the data when it is tested on the test dataset. The function does what it wants as if it was not trained on the data. Parametric models tend to suffer from high bias.
Variance is the change in the function if it was estimated using new data. Variance is usually much higher in non-parametric models as they are more sensitive to the unique nature of each dataset.
Accuracy vs Interpretability
It is also important to determine what you want to know. If your goal is accuracy then a complicated model may be more appropriate. However, if you want to infer and make conclusions about your model then it is preferred to make a simpler model that can be explained.
A model can be made simpler or more complex through the inclusion of more features or the use of a more complex algorithm. For example, regression is much easier to interpret than artificial neural networks.
Supervised vs Unsupervised
Supervised learning is learning that involves a dependent variable. Examples of supervised learning include regression, support vector machines, K nearest neighbor, and random forest.
Unsupervised learning involves a dataset that does not have a dependent variable. In this situation, you are looking for patterns in the data. Examples of this include kmeans, principal component analysis, and cluster analysis.
Regression vs Classifications
Lastly, a problem can call for regression or classification. A regression problem involves a numeric dependent variable. A classification problem has a categorical dependent variable. Almost all models that are used for supervised machine learning can address regression or classification.
For example, regression includes numeric regression and logistic regression for classification. K nearest neighbor can do both as can random forest, support vector machines, and artificial neural networks.

Statistical Learning
Statistical learning is a discipline that focuses on understanding data. Understanding data can happen through classifying or making a numeric prediction which is called supervised learning or finding patterns in data which is called unsupervised learning,
In this post, we will examine the following
- History of statistical learning
- The purpose of statistical learning
- Statistical learning vs Machine learning
History Of Statistical Learning
The early pioneers of statistical learning focused exclusively on supervised learning. Linear regression was developed in the 19th century by Legendre and Gauss. In the 1930’s, Fisher created linear discriminant analysis. Logistic regression was created in the 1940’s as an alternative the linear discriminant analysis.
The developments of the late 19th century to the mid 20th century were limited due to the lack of computational power. However, by the 1970’s things began to change and new algorithms emerged, specifically ones that can handle non-linear relationships
In the 1980’s Friedman and Stone developed classification and regression trees. The term generalized additive models were first used by Hastie and Tibshirani for non-linear generalized models.
Purpose of Statistical Learning
The primary goal of statistical learning is to develop a model of data you currently have to make decisions about the future. In terms of supervised learning with a numeric dependent variable, a teacher may have data on their students and want to predict future academic performance. For a categorical variable, a doctor may use data he has to predict whether someone has cancer or not. In both situations, the goal is to use what one knows to predict what one does not know.
A unique characteristic of supervised learning is that the purpose can be to predict future values or to explain the relationship between the dependent variable and another independent variable(s). Generally, data science is much more focused on prediction while the social sciences seem more concerned with explanations.
For unsupervised learning, there is no dependent variable. In terms of a practical example, a company may want to use the data they have to determine several unique categories of customers they have. Understanding large groups of customer behavior can allow the company to adjust their marketing strategy to cater to the different needs of their vast clientele.
Statistical Learning vs Machine Learning
The difference between statistical learning and machine learning is so small that for the average person it makes little difference. Generally, although some may disagree, these two terms mean essentially the same thing. Often statisticians speak of statistical learning while computer scientist speak of machine learning
Machine learning is the more popular term as it is easier to conceive of a machine learning rather than statistics learning.
Conclusion
Statistical or machine learning is a major force in the world today. With some much data and so much computing power, the possibilities are endless in terms of what kind of beneficial information can be gleaned. However, all this began with people creating a simple linear model in the 19th century.
Linear Regression vs Bayesian Regression
In this post, we are going to look at Bayesian regression. In particular, we will compare the results of ordinary least squares regression with Bayesian regression.
Bayesian Statistics
Bayesian statistics involves the use of probabilities rather than frequencies when addressing uncertainty. This allows you to determine the distribution of the model parameters and not only the values. This is done through averaging over the model parameters through marginalizing the joint probability distribution.
Linear Regression
We will now develop our two models. The first model will be a normal regression and the second a Bayesian model. We will be looking at factors that affect the tax rate of homes in the “Hedonic” dataset in the “Ecdat” package. We will load our packages and partition our data. Below is some initial code
library(ISLR);library(caret);library(arm);library(Ecdat);library(gridExtra)data("Hedonic")
inTrain<-createDataPartition(y=Hedonic$tax,p=0.7, list=FALSE)
trainingset <- Hedonic[inTrain, ]
testingset <- Hedonic[-inTrain, ]
str(Hedonic)## 'data.frame': 506 obs. of 15 variables:
## $ mv : num 10.09 9.98 10.45 10.42 10.5 ...
## $ crim : num 0.00632 0.02731 0.0273 0.03237 0.06905 ...
## $ zn : num 18 0 0 0 0 0 12.5 12.5 12.5 12.5 ...
## $ indus : num 2.31 7.07 7.07 2.18 2.18 ...
## $ chas : Factor w/ 2 levels "no","yes": 1 1 1 1 1 1 1 1 1 1 ...
## $ nox : num 28.9 22 22 21 21 ...
## $ rm : num 43.2 41.2 51.6 49 51.1 ...
## $ age : num 65.2 78.9 61.1 45.8 54.2 ...
## $ dis : num 1.41 1.6 1.6 1.8 1.8 ...
## $ rad : num 0 0.693 0.693 1.099 1.099 ...
## $ tax : int 296 242 242 222 222 222 311 311 311 311 ...
## $ ptratio: num 15.3 17.8 17.8 18.7 18.7 ...
## $ blacks : num 0.397 0.397 0.393 0.395 0.397 ...
## $ lstat : num -3 -2.39 -3.21 -3.53 -2.93 ...
## $ townid : int 1 2 2 3 3 3 4 4 4 4 ...We will now create our regression model
ols.reg<-lm(tax~.,trainingset)
summary(ols.reg)##
## Call:
## lm(formula = tax ~ ., data = trainingset)
##
## Residuals:
## Min 1Q Median 3Q Max
## -180.898 -35.276 2.731 33.574 200.308
##
## Coefficients:
## Estimate Std. Error t value Pr(>|t|)
## (Intercept) 305.1928 192.3024 1.587 0.11343
## mv -41.8746 18.8490 -2.222 0.02697 *
## crim 0.3068 0.6068 0.506 0.61339
## zn 1.3278 0.2006 6.618 1.42e-10 ***
## indus 7.0685 0.8786 8.045 1.44e-14 ***
## chasyes -17.0506 15.1883 -1.123 0.26239
## nox 0.7005 0.4797 1.460 0.14518
## rm -0.1840 0.5875 -0.313 0.75431
## age 0.3054 0.2265 1.349 0.17831
## dis -7.4484 14.4654 -0.515 0.60695
## rad 98.9580 6.0964 16.232 < 2e-16 ***
## ptratio 6.8961 2.1657 3.184 0.00158 **
## blacks -29.6389 45.0043 -0.659 0.51061
## lstat -18.6637 12.4674 -1.497 0.13532
## townid 1.1142 0.1649 6.758 6.07e-11 ***
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
##
## Residual standard error: 63.72 on 341 degrees of freedom
## Multiple R-squared: 0.8653, Adjusted R-squared: 0.8597
## F-statistic: 156.4 on 14 and 341 DF, p-value: < 2.2e-16The model does a reasonable job. Next, we will do our prediction and compare the results with the test set using correlation, summary statistics, and the mean absolute error. In the code below, we use the “predict.lm” function and include the arguments “interval” for the prediction as well as “se.fit” for the standard error
ols.regTest<-predict.lm(ols.reg,testingset,interval = 'prediction',se.fit = T)Below is the code for the correlation, summary stats, and mean absolute error. For MAE, we need to create a function.
cor(testingset$tax,ols.regTest$fit[,1])## [1] 0.9313795summary(ols.regTest$fit[,1])## Min. 1st Qu. Median Mean 3rd Qu. Max.
## 144.7 288.3 347.6 399.4 518.4 684.1summary(trainingset$tax)## Min. 1st Qu. Median Mean 3rd Qu. Max.
## 188.0 279.0 330.0 410.4 666.0 711.0MAE<-function(actual, predicted){
mean(abs(actual-predicted))
}
MAE(ols.regTest$fit[,1], testingset$tax)## [1] 41.07212The correlation is excellent. The summary stats are similar and the error is not unreasonable. Below is a plot of the actual and predicted values
We now need to combine some data into one dataframe. In particular, we need the following actual dependent variable results predicted dependent variable results The upper confidence value of the prediction THe lower confidence value of the prediction
The code is below
yout.ols <- as.data.frame(cbind(testingset$tax,ols.regTest$fit))
ols.upr <- yout.ols$upr
ols.lwr <- yout.ols$lwrWe can now plot this
p.ols <- ggplot(data = yout.ols, aes(x = testingset$tax, y = ols.regTest$fit[,1])) + geom_point() + ggtitle("Ordinary Regression") + labs(x = "Actual", y = "Predicted")
p.ols + geom_errorbar(ymin = ols.lwr, ymax = ols.upr)
You can see the strong linear relationship. However, the confidence intervals are rather wide. Let’s see how Bayes does.
Bayes Regression
Bayes regression uses the “bayesglm” function from the “arm” package. We need to set the family to “gaussian” and the link to “identity”. In addition, we have to set the “prior.df” (prior degrees of freedom) to infinity as this indicates we want a normal distribution
bayes.reg<-bayesglm(tax~.,family=gaussian(link=identity),trainingset,prior.df = Inf)bayes.regTest<-predict.glm(bayes.reg,newdata = testingset,se.fit = T)
cor(testingset$tax,bayes.regTest$fit)## [1] 0.9313793summary(bayes.regTest$fit)## Min. 1st Qu. Median Mean 3rd Qu. Max.
## 144.7 288.3 347.5 399.4 518.4 684.1summary(trainingset$tax)## Min. 1st Qu. Median Mean 3rd Qu. Max.
## 188.0 279.0 330.0 410.4 666.0 711.0MAE(bayes.regTest$fit, testingset$tax)## [1] 41.07352The numbers are essentially the same. This leads to the question of what is the benefit of Bayesian regression? The answer is in the confidence intervals. Next, we will calculate the confidence intervals for the Bayesian model.
yout.bayes <- as.data.frame(cbind(testingset$tax,bayes.regTest$fit))
names(yout.bayes) <- c("tax", "fit")
critval <- 1.96 #approx for 95% CI
bayes.upr <- bayes.regTest$fit + critval * bayes.regTest$se.fit
bayes.lwr <- bayes.regTest$fit - critval * bayes.regTest$se.fitWe now create our Bayesian regression plot.
p.bayes <- ggplot(data = yout.bayes, aes(x = yout.bayes$tax, y = yout.bayes$fit)) + geom_point() + ggtitle("Bayesian Regression Prediction") + labs(x = "Actual", y = "Predicted")Lastly, we display both plots as a comparison.
ols.plot <- p.ols + geom_errorbar(ymin = ols.lwr, ymax = ols.upr)
bayes.plot <- p.bayes + geom_errorbar(ymin = bayes.lwr, ymax = bayes.upr)
grid.arrange(ols.plot,bayes.plot,ncol=2)
As you can see, the Bayesian approach gives much more compact confidence intervals. This is because the Bayesian approach a distribution of parameters is calculated from a posterior distribution. These values are then averaged to get the final prediction that appears on the plot. This reduces the variance and strengthens the confidence we can have in each individual example.
Statistical Models
In research, the term ‘model’ is employed frequently. Normally, a model is some sort of a description or explanation of a real world phenomenon. In data science, we employ the use of statistical models. Statistical models used numbers to help us to understand something that happens in the real world.
A statistical model used numbers to help us to understand something that happens in the real world.
In the real world, quantitative research relies on numeric observations of some phenomenon, behavior, and or perception. For example, let’s say we have the quiz results of 20 students as show below.
32 60 95 15 43 22 45 14 48 98 79 97 49 63 50 11 26 52 39 97
This is great information but what if we want to go beyond how these students did and try to understand how students in the population would do on the quizzes. Doing this requires the development of a model.
A model is simply trying to describe how the data is generated in terms of whatever we are mesuring while allowing for randomness. It helps in summarizing a large collection of numbers while also providing structure to it.
One commonly used model is the normal model. This model is the famous bell-curve model that most of us are familiar with. To calculate this model we need to calculate the mean and standard deviation to get a plot similar to the one below

Now, this model is not completely perfect. For example, a student cannot normally get a score above 100 or below 0 on a quiz. Despite this weakness, normal distribution gives is an indication of what the population looks like.
With this, we can also calculate the probability of getting a specific score on the quiz. For example, if we want to calculate the probability that a student would get a score of 70 or higher we can do a simple test and find that it is about 26%.
Other Options
The normal model is not the only model. There are many different models to match different types of data. There are the gamma, student t, binomial, chi-square, etc. To determine which model to use requires examining the distribution of your data and match it to an appropriate model.
Another option is to transform the data. This is normally done to make data conform to a normal distribution. Which transformation to employ depends on how the data looks when it is plotted.
Conclusion
Modeling helps to bring order to data that has been collected for analysis. By using a model such as the normal distribution, you can begin to make inferences about what the population is like. This allows you to take a handful of data to better understand the world.
K Nearest Neighbor in R
K-nearest neighbor is one of many nonlinear algorithms that can be used in machine learning. By non-linear I mean that a linear combination of the features or variables is not needed in order to develop decision boundaries. This allows for the analysis of data that naturally does not meet the assumptions of linearity.
KNN is also known as a “lazy learner”. This means that there are known coefficients or parameter estimates. When doing regression we always had coefficient outputs regardless of the type of regression (ridge, lasso, elastic net, etc.). What KNN does instead is used K nearest neighbors to give a label to an unlabeled example. Our job when using KNN is to determine the number of K neighbors to use that is most accurate based on the different criteria for assessing the models.
In this post, we will develop a KNN model using the “Mroz” dataset from the “Ecdat” package. Our goal is to predict if someone lives in the city based on the other predictor variables. Below is some initial code.
library(class);library(kknn);library(caret);library(corrplot)library(reshape2);library(ggplot2);library(pROC);library(Ecdat)
data(Mroz) str(Mroz)
## 'data.frame': 753 obs. of 18 variables:
## $ work : Factor w/ 2 levels "yes","no": 2 2 2 2 2 2 2 2 2 2 ...
## $ hoursw : int 1610 1656 1980 456 1568 2032 1440 1020 1458 1600 ...
## $ child6 : int 1 0 1 0 1 0 0 0 0 0 ...
## $ child618 : int 0 2 3 3 2 0 2 0 2 2 ...
## $ agew : int 32 30 35 34 31 54 37 54 48 39 ...
## $ educw : int 12 12 12 12 14 12 16 12 12 12 ...
## $ hearnw : num 3.35 1.39 4.55 1.1 4.59 ...
## $ wagew : num 2.65 2.65 4.04 3.25 3.6 4.7 5.95 9.98 0 4.15 ...
## $ hoursh : int 2708 2310 3072 1920 2000 1040 2670 4120 1995 2100 ...
## $ ageh : int 34 30 40 53 32 57 37 53 52 43 ...
## $ educh : int 12 9 12 10 12 11 12 8 4 12 ...
## $ wageh : num 4.03 8.44 3.58 3.54 10 ...
## $ income : int 16310 21800 21040 7300 27300 19495 21152 18900 20405 20425 ...
## $ educwm : int 12 7 12 7 12 14 14 3 7 7 ...
## $ educwf : int 7 7 7 7 14 7 7 3 7 7 ...
## $ unemprate : num 5 11 5 5 9.5 7.5 5 5 3 5 ...
## $ city : Factor w/ 2 levels "no","yes": 1 2 1 1 2 2 1 1 1 1 ...
## $ experience: int 14 5 15 6 7 33 11 35 24 21 ...We need to remove the factor variable “work” as KNN cannot use factor variables. After this, we will use the “melt” function from the “reshape2” package to look at the variables when divided by whether the example was from the city or not.
Mroz$work<-NULL
mroz.melt<-melt(Mroz,id.var='city')
Mroz_plots<-ggplot(mroz.melt,aes(x=city,y=value))+geom_boxplot()+facet_wrap(~variable, ncol = 4)
Mroz_plots
From the plots, it appears there are no differences in how the variable act whether someone is from the city or not. This may be a flag that classification may not work.
We now need to scale our data otherwise the results will be inaccurate. Scaling might also help our box-plots because everything will be on the same scale rather than spread all over the place. To do this we will have to temporarily remove our outcome variable from the data set because it’s a factor and then reinsert it into the data set. Below is the code.
mroz.scale<-as.data.frame(scale(Mroz[,-16]))
mroz.scale$city<-Mroz$cityWe will now look at our box-plots a second time but this time with scaled data.
mroz.scale.melt<-melt(mroz.scale,id.var="city")
mroz_plot2<-ggplot(mroz.scale.melt,aes(city,value))+geom_boxplot()+facet_wrap(~variable, ncol = 4)
mroz_plot2
This second plot is easier to read but there is still little indication of difference.
We can now move to checking the correlations among the variables. Below is the code
mroz.cor<-cor(mroz.scale[,-17])
corrplot(mroz.cor,method = 'number')
There is a high correlation between husband’s age (ageh) and wife’s age (agew). Since this algorithm is non-linear this should not be a major problem.
We will now divide our dataset into the training and testing sets
set.seed(502)
ind=sample(2,nrow(mroz.scale),replace=T,prob=c(.7,.3))
train<-mroz.scale[ind==1,]
test<-mroz.scale[ind==2,]Before creating a model we need to create a grid. We do not know the value of k yet so we have to run multiple models with different values of k in order to determine this for our model. As such we need to create a ‘grid’ using the ‘expand.grid’ function. We will also use cross-validation to get a better estimate of k as well using the “trainControl” function. The code is below.
grid<-expand.grid(.k=seq(2,20,by=1))
control<-trainControl(method="cv")Now we make our model,
knn.train<-train(city~.,train,method="knn",trControl=control,tuneGrid=grid)
knn.train## k-Nearest Neighbors
##
## 540 samples
## 16 predictors
## 2 classes: 'no', 'yes'
##
## No pre-processing
## Resampling: Cross-Validated (10 fold)
## Summary of sample sizes: 487, 486, 486, 486, 486, 486, ...
## Resampling results across tuning parameters:
##
## k Accuracy Kappa
## 2 0.6000095 0.1213920
## 3 0.6368757 0.1542968
## 4 0.6424325 0.1546494
## 5 0.6386252 0.1275248
## 6 0.6329998 0.1164253
## 7 0.6589619 0.1616377
## 8 0.6663344 0.1774391
## 9 0.6663681 0.1733197
## 10 0.6609510 0.1566064
## 11 0.6664018 0.1575868
## 12 0.6682199 0.1669053
## 13 0.6572111 0.1397222
## 14 0.6719586 0.1694953
## 15 0.6571425 0.1263937
## 16 0.6664367 0.1551023
## 17 0.6719573 0.1588789
## 18 0.6608811 0.1260452
## 19 0.6590979 0.1165734
## 20 0.6609510 0.1219624
##
## Accuracy was used to select the optimal model using the largest value.
## The final value used for the model was k = 14.R recommends that k = 16. This is based on a combination of accuracy and the kappa statistic. The kappa statistic is a measurement of the accuracy of a model while taking into account chance. We don’t have a model in the sense that we do not use the ~ sign like we do with regression. Instead, we have a train and a test set a factor variable and a number for k. This will make more sense when you see the code. Finally, we will use this information on our test dataset. We will then look at the table and the accuracy of the model.
knn.test<-knn(train[,-17],test[,-17],train[,17],k=16) #-17 removes the dependent variable 'city
table(knn.test,test$city)##
## knn.test no yes
## no 19 8
## yes 61 125prob.agree<-(15+129)/213
prob.agree## [1] 0.6760563Accuracy is 67% which is consistent with what we found when determining the k. We can also calculate the kappa. This done by calculating the probability and then do some subtraction and division. We already know the accuracy as we stored it in the variable “prob.agree” we now need the probability that this is by chance. Lastly, we calculate the kappa.
prob.chance<-((15+4)/213)*((15+65)/213)
kap<-(prob.agree-prob.chance)/(1-prob.chance)
kap## [1] 0.664827A kappa of .66 is actually good.
The example we just did was with unweighted k neighbors. There are times when weighted neighbors can improve accuracy. We will look at three different weighing methods. “Rectangular” is unweighted and is the one that we used. The other two are “triangular” and “epanechnikov”. How these calculate the weights is beyond the scope of this post. In the code below the argument “distance” can be set to 1 for Euclidean and 2 for absolute distance.
kknn.train<-train.kknn(city~.,train,kmax = 25,distance = 2,kernel = c("rectangular","triangular",
"epanechnikov"))
plot(kknn.train)
kknn.train##
## Call:
## train.kknn(formula = city ~ ., data = train, kmax = 25, distance = 2, kernel = c("rectangular", "triangular", "epanechnikov"))
##
## Type of response variable: nominal
## Minimal misclassification: 0.3277778
## Best kernel: rectangular
## Best k: 14If you look at the plot you can see which value of k is the best by looking at the point that is the lowest on the graph which is right before 15. Looking at the legend it indicates that the point is the “rectangular” estimate which is the same as unweighted. This means that the best classification is unweighted with a k of 14. Although it recommends a different value for k our misclassification was about the same.
Conclusion
In this post, we explored both weighted and unweighted KNN. This algorithm allows you to deal with data that does not meet the assumptions of regression by ignoring the need for parameters. However, because there are no numbers really attached to the results beyond accuracy it can be difficult to explain what is happening in the model to people. As such, perhaps the biggest drawback is communicating results when using KNN.
Elastic Net Regression in R
Elastic net is a combination of ridge and lasso regression. What is most unusual about elastic net is that it has two tuning parameters (alpha and lambda) while lasso and ridge regression only has 1.
In this post, we will go through an example of the use of elastic net using the “VietnamI” dataset from the “Ecdat” package. Our goal is to predict how many days a person is ill based on the other variables in the dataset. Below is some initial code for our analysis
library(Ecdat);library(corrplot);library(caret);library(glmnet)data("VietNamI") str(VietNamI)
## 'data.frame': 27765 obs. of 12 variables:
## $ pharvis : num 0 0 0 1 1 0 0 0 2 3 ...
## $ lnhhexp : num 2.73 2.74 2.27 2.39 3.11 ...
## $ age : num 3.76 2.94 2.56 3.64 3.3 ...
## $ sex : Factor w/ 2 levels "female","male": 2 1 2 1 2 2 1 2 1 2 ...
## $ married : num 1 0 0 1 1 1 1 0 1 1 ...
## $ educ : num 2 0 4 3 3 9 2 5 2 0 ...
## $ illness : num 1 1 0 1 1 0 0 0 2 1 ...
## $ injury : num 0 0 0 0 0 0 0 0 0 0 ...
## $ illdays : num 7 4 0 3 10 0 0 0 4 7 ...
## $ actdays : num 0 0 0 0 0 0 0 0 0 0 ...
## $ insurance: num 0 0 1 1 0 1 1 1 0 0 ...
## $ commune : num 192 167 76 123 148 20 40 57 49 170 ...
## - attr(*, "na.action")=Class 'omit' Named int 27734
## .. ..- attr(*, "names")= chr "27734"We need to check the correlations among the variables. We need to exclude the “sex” variable as it is categorical. The code is below.
p.cor<-cor(VietNamI[,-4])
corrplot.mixed(p.cor)
No major problems with correlations. Next, we set up our training and testing datasets. We need to remove the variable “commune” because it adds no value to our results. In addition, to reduce the computational time we will only use the first 1000 rows from the data set.
VietNamI$commune<-NULL
VietNamI_reduced<-VietNamI[1:1000,]
ind<-sample(2,nrow(VietNamI_reduced),replace=T,prob = c(0.7,0.3))
train<-VietNamI_reduced[ind==1,]
test<-VietNamI_reduced[ind==2,]We need to create a grid that will allow us to investigate different models with different combinations of alpha and lambda. This is done using the “expand.grid” function. In combination with the “seq” function below is the code
grid<-expand.grid(.alpha=seq(0,1,by=.5),.lambda=seq(0,0.2,by=.1))We also need to set the resampling method, which allows us to assess the validity of our model. This is done using the “trainControl” function” from the “caret” package. In the code below “LOOCV” stands for “leave one out cross-validation”.
control<-trainControl(method = "LOOCV")We are no ready to develop our model. The code is mostly self-explanatory. This initial model will help us to determine the appropriate values for the alpha and lambda parameters
enet.train<-train(illdays~.,train,method="glmnet",trControl=control,tuneGrid=grid)
enet.train## glmnet
##
## 694 samples
## 10 predictors
##
## No pre-processing
## Resampling: Leave-One-Out Cross-Validation
## Summary of sample sizes: 693, 693, 693, 693, 693, 693, ...
## Resampling results across tuning parameters:
##
## alpha lambda RMSE Rsquared
## 0.0 0.0 5.229759 0.2968354
## 0.0 0.1 5.229759 0.2968354
## 0.0 0.2 5.229759 0.2968354
## 0.5 0.0 5.243919 0.2954226
## 0.5 0.1 5.225067 0.2985989
## 0.5 0.2 5.200415 0.3038821
## 1.0 0.0 5.244020 0.2954519
## 1.0 0.1 5.203973 0.3033173
## 1.0 0.2 5.182120 0.3083819
##
## RMSE was used to select the optimal model using the smallest value.
## The final values used for the model were alpha = 1 and lambda = 0.2.The output list all the possible alpha and lambda values that we set in the “grid” variable. It even tells us which combination was the best. For our purposes, the alpha will be .5 and the lambda .2. The r-square is also included.
We will set our model and run it on the test set. We have to convert the “sex” variable to a dummy variable for the “glmnet” function. We next have to make matrices for the predictor variables and a for our outcome variable “illdays”
train$sex<-model.matrix( ~ sex - 1, data=train ) #convert to dummy variable
test$sex<-model.matrix( ~ sex - 1, data=test )
predictor_variables<-as.matrix(train[,-9])
days_ill<-as.matrix(train$illdays)
enet<-glmnet(predictor_variables,days_ill,family = "gaussian",alpha = 0.5,lambda = .2)We can now look at specific coefficient by using the “coef” function.
enet.coef<-coef(enet,lambda=.2,alpha=.5,exact=T)
enet.coef## 12 x 1 sparse Matrix of class "dgCMatrix"
## s0
## (Intercept) -1.304263895
## pharvis 0.532353361
## lnhhexp -0.064754000
## age 0.760864404
## sex.sexfemale 0.029612290
## sex.sexmale -0.002617404
## married 0.318639271
## educ .
## illness 3.103047473
## injury .
## actdays 0.314851347
## insurance .You can see for yourself that several variables were removed from the model. Medical expenses (lnhhexp), sex, education, injury, and insurance do not play a role in the number of days ill for an individual in Vietnam.
With our model developed. We now can test it using the predict function. However, we first need to convert our test dataframe into a matrix and remove the outcome variable from it
test.matrix<-as.matrix(test[,-9])
enet.y<-predict(enet, newx = test.matrix, type = "response", lambda=.2,alpha=.5)Let’s plot our results
plot(enet.y)
This does not look good. Let’s check the mean squared error
enet.resid<-enet.y-test$illdays
mean(enet.resid^2)## [1] 20.18134We will now do a cross-validation of our model. We need to set the seed and then use the “cv.glmnet” to develop the cross-validated model. We can see the model by plotting it.
set.seed(317)
enet.cv<-cv.glmnet(predictor_variables,days_ill,alpha=.5)
plot(enet.cv)
You can see that as the number of features are reduce (see the numbers on the top of the plot) the MSE increases (y-axis). In addition, as the lambda increases, there is also an increase in the error but only when the number of variables is reduced as well.
The dotted vertical lines in the plot represent the minimum MSE for a set lambda (on the left) and the one standard error from the minimum (on the right). You can extract these two lambda values using the code below.
enet.cv$lambda.min## [1] 0.3082347enet.cv$lambda.1se## [1] 2.874607We can see the coefficients for a lambda that is one standard error away by using the code below. This will give us an alternative idea for what to set the model parameters to when we want to predict.
coef(enet.cv,s="lambda.1se")## 12 x 1 sparse Matrix of class "dgCMatrix"
## 1
## (Intercept) 2.34116947
## pharvis 0.003710399
## lnhhexp .
## age .
## sex.sexfemale .
## sex.sexmale .
## married .
## educ .
## illness 1.817479480
## injury .
## actdays .
## insurance .Using the one standard error lambda we lose most of our features. We can now see if the model improves by rerunning it with this information.
enet.y.cv<-predict(enet.cv,newx = test.matrix,type='response',lambda="lambda.1se", alpha = .5)
enet.cv.resid<-enet.y.cv-test$illdays
mean(enet.cv.resid^2)## [1] 25.47966A small improvement. Our model is a mess but this post served as an example of how to conduct an analysis using elastic net regression.
Lasso Regression in R
In this post, we will conduct an analysis using the lasso regression. Remember lasso regression will actually eliminate variables by reducing them to zero through how the shrinkage penalty can be applied.
We will use the dataset “nlschools” from the “MASS” packages to conduct our analysis. We want to see if we can predict language test scores “lang” with the other available variables. Below is some initial code to begin the analysis
library(MASS);library(corrplot);library(glmnet)data("nlschools") str(nlschools)
## 'data.frame': 2287 obs. of 6 variables:
## $ lang : int 46 45 33 46 20 30 30 57 36 36 ...
## $ IQ : num 15 14.5 9.5 11 8 9.5 9.5 13 9.5 11 ...
## $ class: Factor w/ 133 levels "180","280","1082",..: 1 1 1 1 1 1 1 1 1 1 ...
## $ GS : int 29 29 29 29 29 29 29 29 29 29 ...
## $ SES : int 23 10 15 23 10 10 23 10 13 15 ...
## $ COMB : Factor w/ 2 levels "0","1": 1 1 1 1 1 1 1 1 1 1 ...We need to remove the “class” variable as it is used as an identifier and provides no useful data. After this, we can check the correlations among the variables. Below is the code for this.
nlschools$class<-NULL
p.cor<-cor(nlschools[,-5])
corrplot.mixed(p.cor)
No problems with collinearity. We will now setup are training and testing sets.
ind<-sample(2,nrow(nlschools),replace=T,prob = c(0.7,0.3))
train<-nlschools[ind==1,]
test<-nlschools[ind==2,]Remember that the ‘glmnet’ function does not like factor variables. So we need to convert our “COMB” variable to a dummy variable. In addition, “glmnet” function does not like data frames so we need to make two data frames. The first will include all the predictor variables and the second we include only the outcome variable. Below is the code
train$COMB<-model.matrix( ~ COMB - 1, data=train ) #convert to dummy variable
test$COMB<-model.matrix( ~ COMB - 1, data=test )
predictor_variables<-as.matrix(train[,2:4])
language_score<-as.matrix(train$lang)We can now run our model. We place both matrices inside the “glmnet” function. The family is set to “gaussian” because our outcome variable is continuous. The “alpha” is set to 1 as this indicates that we are using lasso regression.
lasso<-glmnet(predictor_variables,language_score,family="gaussian",alpha=1)Now we need to look at the results using the “print” function. This function prints a lot of information as explained below.
- Df = number of variables including in the model (this is always the same number in a ridge model)
- %Dev = Percent of deviance explained. The higher the better
- Lambda = The lambda used to obtain the %Dev
When you use the “print” function for a lasso model it will print up to 100 different models. Fewer models are possible if the percent of deviance stops improving. 100 is the default stopping point. In the code below we will use the “print” function but, I only printed the first 5 and last 5 models in order to reduce the size of the printout. Fortunately, it only took 60 models to converge.
print(lasso)##
## Call: glmnet(x = predictor_variables, y = language_score, family = "gaussian", alpha = 1)
##
## Df %Dev Lambda
## [1,] 0 0.00000 5.47100
## [2,] 1 0.06194 4.98500
## [3,] 1 0.11340 4.54200
## [4,] 1 0.15610 4.13900
## [5,] 1 0.19150 3.77100
............................
## [55,] 3 0.39890 0.03599
## [56,] 3 0.39900 0.03280
## [57,] 3 0.39900 0.02988
## [58,] 3 0.39900 0.02723
## [59,] 3 0.39900 0.02481
## [60,] 3 0.39900 0.02261The results from the “print” function will allow us to set the lambda for the “test” dataset. Based on the results we can set the lambda at 0.02 because this explains the highest amount of deviance at .39.
The plot below shows us lambda on the x-axis and the coefficients of the predictor variables on the y-axis. The numbers next to the coefficient lines refers to the actual coefficient of a particular variable as it changes from using different lambda values. Each number corresponds to a variable going from left to right in a dataframe/matrix using the “View” function. For example, 1 in the plot refers to “IQ” 2 refers to “GS” etc.
plot(lasso,xvar="lambda",label=T)
As you can see, as lambda increase the coefficient decrease in value. This is how regularized regression works. However, unlike ridge regression which never reduces a coefficient to zero, lasso regression does reduce a coefficient to zero. For example, coefficient 3 (SES variable) and coefficient 2 (GS variable) are reduced to zero when lambda is near 1.
You can also look at the coefficient values at a specific lambda values. The values are unstandardized and are used to determine the final model selection. In the code below the lambda is set to .02 and we use the “coef” function to do see the results
lasso.coef<-coef(lasso,s=.02,exact = T)
lasso.coef## 4 x 1 sparse Matrix of class "dgCMatrix"
## 1
## (Intercept) 9.35736325
## IQ 2.34973922
## GS -0.02766978
## SES 0.16150542Results indicate that for a 1 unit increase in IQ there is a 2.41 point increase in language score. When GS (class size) goes up 1 unit there is a .03 point decrease in language score. Finally, when SES (socioeconomic status) increase 1 unit language score improves .13 point.
The second plot shows us the deviance explained on the x-axis. On the y-axis is the coefficients of the predictor variables. Below is the code
plot(lasso,xvar='dev',label=T)
If you look carefully, you can see that the two plots are completely opposite to each other. increasing lambda cause a decrease in the coefficients. Furthermore, increasing the fraction of deviance explained leads to an increase in the coefficient. You may remember seeing this when we used the “print”” function. As lambda became smaller there was an increase in the deviance explained.
Now, we will assess our model using the test data. We need to convert the test dataset to a matrix. Then we will use the “predict”” function while setting our lambda to .02. Lastly, we will plot the results. Below is the code.
test.matrix<-as.matrix(test[,2:4])
lasso.y<-predict(lasso,newx = test.matrix,type = 'response',s=.02)
plot(lasso.y,test$lang)
The visual looks promising. The last thing we need to do is calculated the mean squared error. By its self this number does not mean much. However, it provides a benchmark for comparing our current model with any other models that we may develop. Below is the code
lasso.resid<-lasso.y-test$lang
mean(lasso.resid^2)## [1] 46.74314Knowing this number, we can, if we wanted, develop other models using other methods of analysis to try to reduce it. Generally, the lower the error the better while keeping in mind the complexity of the model.
Ridge Regression in R
In this post, we will conduct an analysis using ridge regression. Ridge regression is a type of regularized regression. By applying a shrinkage penalty, we are able to reduce the coefficients of many variables almost to zero while still retaining them in the model. This allows us to develop models that have many more variables in them compared to models using the best subset or stepwise regression.
In the example used in this post, we will use the “SAheart” dataset from the “ElemStatLearn” package. We want to predict systolic blood pressure (sbp) using all of the other variables available as predictors. Below is some initial code that we need to begin.
library(ElemStatLearn);library(car);library(corrplot)
library(leaps);library(glmnet);library(caret)data(SAheart)
str(SAheart)
## 'data.frame': 462 obs. of 10 variables:
## $ sbp : int 160 144 118 170 134 132 142 114 114 132 ...
## $ tobacco : num 12 0.01 0.08 7.5 13.6 6.2 4.05 4.08 0 0 ...
## $ ldl : num 5.73 4.41 3.48 6.41 3.5 6.47 3.38 4.59 3.83 5.8 ...
## $ adiposity: num 23.1 28.6 32.3 38 27.8 ...
## $ famhist : Factor w/ 2 levels "Absent","Present": 2 1 2 2 2 2 1 2 2 2 ...
## $ typea : int 49 55 52 51 60 62 59 62 49 69 ...
## $ obesity : num 25.3 28.9 29.1 32 26 ...
## $ alcohol : num 97.2 2.06 3.81 24.26 57.34 ...
## $ age : int 52 63 46 58 49 45 38 58 29 53 ...
## $ chd : int 1 1 0 1 1 0 0 1 0 1 ...A look at the object using the “str” function indicates that one variable “famhist” is a factor variable. The “glmnet” function that does the ridge regression analysis cannot handle factors so we need to convert this to a dummy variable. However, there are two things we need to do before this. First, we need to check the correlations to make sure there are no major issues with multicollinearity Second, we need to create our training and testing data sets. Below is the code for the correlation plot.
p.cor<-cor(SAheart[,-5])
corrplot.mixed(p.cor)
First we created a variable called “p.cor” the -5 in brackets means we removed the 5th column from the “SAheart” data set which is the factor variable “Famhist”. The correlation plot indicates that there is one strong relationship between adiposity and obesity. However, one common cut-off for collinearity is 0.8 and this value is 0.72 which is not a problem.
We will now create are training and testing sets and convert “famhist” to a dummy variable.
ind<-sample(2,nrow(SAheart),replace=T,prob = c(0.7,0.3))
train<-SAheart[ind==1,]
test<-SAheart[ind==2,]
train$famhist<-model.matrix( ~ famhist - 1, data=train ) #convert to dummy variable
test$famhist<-model.matrix( ~ famhist - 1, data=test )We are still not done preparing our data yet. “glmnet” cannot use data frames, instead, it can only use matrices. Therefore, we now need to convert our data frames to matrices. We have to create two matrices, one with all of the predictor variables and a second with the outcome variable of blood pressure. Below is the code
predictor_variables<-as.matrix(train[,2:10])
blood_pressure<-as.matrix(train$sbp)We are now ready to create our model. We use the “glmnet” function and insert our two matrices. The family is set to Gaussian because “blood pressure” is a continuous variable. Alpha is set to 0 as this indicates ridge regression. Below is the code
ridge<-glmnet(predictor_variables,blood_pressure,family = 'gaussian',alpha = 0)Now we need to look at the results using the “print” function. This function prints a lot of information as explained below.
- Df = number of variables including in the model (this is always the same number in a ridge model)
- %Dev = Percent of deviance explained. The higher the better
- Lambda = The lambda used to attain the %Dev
When you use the “print” function for a ridge model it will print up to 100 different models. Fewer models are possible if the percent of deviance stops improving. 100 is the default stopping point. In the code below we have the “print” function. However, I have only printed the first 5 and last 5 models in order to save space.
print(ridge)##
## Call: glmnet(x = predictor_variables, y = blood_pressure, family = "gaussian", alpha = 0)
##
## Df %Dev Lambda
## [1,] 10 7.622e-37 7716.0000
## [2,] 10 2.135e-03 7030.0000
## [3,] 10 2.341e-03 6406.0000
## [4,] 10 2.566e-03 5837.0000
## [5,] 10 2.812e-03 5318.0000
................................
## [95,] 10 1.690e-01 1.2290
## [96,] 10 1.691e-01 1.1190
## [97,] 10 1.692e-01 1.0200
## [98,] 10 1.693e-01 0.9293
## [99,] 10 1.693e-01 0.8468
## [100,] 10 1.694e-01 0.7716The results from the “print” function are useful in setting the lambda for the “test” dataset. Based on the results we can set the lambda at 0.83 because this explains the highest amount of deviance at .20.
The plot below shows us lambda on the x-axis and the coefficients of the predictor variables on the y-axis. The numbers refer to the actual coefficient of a particular variable. Inside the plot, each number corresponds to a variable going from left to right in a data-frame/matrix using the “View” function. For example, 1 in the plot refers to “tobacco” 2 refers to “ldl” etc. Across the top of the plot is the number of variables used in the model. Remember this number never changes when doing ridge regression.
plot(ridge,xvar="lambda",label=T)
As you can see, as lambda increase the coefficient decrease in value. This is how ridge regression works yet no coefficient ever goes to absolute 0.
You can also look at the coefficient values at a specific lambda value. The values are unstandardized but they provide a useful insight when determining final model selection. In the code below the lambda is set to .83 and we use the “coef” function to do this
ridge.coef<-coef(ridge,s=.83,exact = T)
ridge.coef## 11 x 1 sparse Matrix of class "dgCMatrix"
## 1
## (Intercept) 105.69379942
## tobacco -0.25990747
## ldl -0.13075557
## adiposity 0.29515034
## famhist.famhistAbsent 0.42532887
## famhist.famhistPresent -0.40000846
## typea -0.01799031
## obesity 0.29899976
## alcohol 0.03648850
## age 0.43555450
## chd -0.26539180The second plot shows us the deviance explained on the x-axis and the coefficients of the predictor variables on the y-axis. Below is the code
plot(ridge,xvar='dev',label=T)
The two plots are completely opposite to each other. Increasing lambda cause a decrease in the coefficients while increasing the fraction of deviance explained leads to an increase in the coefficient. You can also see this when we used the “print” function. As lambda became smaller there was an increase in the deviance explained.
We now can begin testing our model on the test data set. We need to convert the test dataset to a matrix and then we will use the predict function while setting our lambda to .83 (remember a lambda of .83 explained the most of the deviance). Lastly, we will plot the results. Below is the code.
test.matrix<-as.matrix(test[,2:10])
ridge.y<-predict(ridge,newx = test.matrix,type = 'response',s=.83)
plot(ridge.y,test$sbp)
The last thing we need to do is calculated the mean squared error. By it’s self this number is useless. However, it provides a benchmark for comparing the current model with any other models you may develop. Below is the code
ridge.resid<-ridge.y-test$sbp
mean(ridge.resid^2)## [1] 372.4431Knowing this number, we can develop other models using other methods of analysis to try to reduce it as much as possible.

Regularized Linear Regression
Traditional linear regression has been a tried and true model for making predictions for decades. However, with the growth of Big Data and datasets with 100’s of variables problems have begun to arise. For example, using stepwise or best subset method with regression could take hours if not days to converge in even some of the best computers.
To deal with this problem, regularized regression has been developed to help to determine which features or variables to keep when developing models from large datasets with a huge number of variables. In this post, we will look at the following concepts
- Definition of regularized regression
- Ridge regression
- Lasso regression
- Elastic net regression
Regularization
Regularization involves the use of a shrinkage penalty in order to reduce the residual sum of squares (RSS). This is done by selecting a value for a tuning parameter called “lambda”. Tuning parameters are used in machine learning algorithms to control the behavior of the models that are developed.
The lambda is multiplied by the normalized coefficients of the model and added to the RSS. Below is an equation of what was just said
RSS + λ(normalized coefficients)
The benefits of regularization are at least three-fold. First, regularization is highly computationally efficient. Instead of fitting k-1 models when k is the number of variables available (for example, 50 variables would lead 49 models!), with regularization only one model is developed for each value of lambda you specify.
Second, regularization helps to deal with the bias-variance headache of model development. When small changes are made to data, such as switching from the training to testing data, there can be wild changes in the estimates. Regularization can often smooth this problem out substantially.
Finally, regularization can help to reduce or eliminate any multicollinearity in a model. As such, the benefits of using regularization make it clear that this should be considered when working with larger datasets.
Ridge Regression
Ridge regression involves the normalization of the squared weights or as shown in the equation below
RSS + λ(normalized coefficients^2)
This is also referred to as the L2-norm. As lambda increase in value, the coefficients in the model are shrunk towards 0 but never reach 0. This is how the error is shrunk. The higher the lambda the lower the value of the coefficients as they are reduced more and more thus reducing the RSS.
The benefit is that predictive accuracy is often increased. However, interpreting and communicating your results can become difficult because no variables are removed from the model. Instead, the variables are reduced near to zero. This can be especially tough if you have dozens of variables remaining in your model to try to explain.
Lasso
Lasso is short for “Least Absolute Shrinkage and Selection Operator”. This approach uses the L1-norm which is the sum of the absolute value of the coefficients or as shown in the equation below
RSS + λ(Σ|normalized coefficients|)
This shrinkage penalty will reduce a coefficient to 0 which is another way of saying that variables will be removed from the model. One problem is that highly correlated variables that need to be in your model may be removed when Lasso shrinks coefficients. This is one reason why ridge regression is still used.
Elastic Net
Elastic net is the best of ridge and Lasso without the weaknesses of either. It combines extracts variables like Lasso and Ridge does not while also group variables like Ridge does but Lasso does not.
This is done by including a second tuning parameter called “alpha”. If alpha is set to 0 it is the same as ridge regression and if alpha is set to 1 it is the same as lasso regression. If you are able to appreciate it below is the formula used for elastic net regression
(RSS + l[(1 – alpha)(S|normalized coefficients|2)/2 + alpha(S|normalized coefficients|)])/N)
As such when working with elastic net you have to set two different tuning parameters (alpha and lambda) in order to develop a model.
Conclusion
Regularized regression was developed as an answer to the growth in the size and number of variables in a data set today. Ridge, lasso an elastic net all provide solutions to converging over large datasets and selecting features.
Logistic Regression in R
In this post, we will conduct a logistic regression analysis. Logistic regression is used when you want to predict a categorical dependent variable using continuous or categorical dependent variables. In our example, we want to predict Sex (male or female) when using several continuous variables from the “survey” dataset in the “MASS” package.
library(MASS);library(bestglm);library(reshape2);library(corrplot)data(survey)
?MASS::survey #explains the variables in the studyThe first thing we need to do is remove the independent factor variables from our dataset. The reason for this is that the function that we will use for the cross-validation does not accept factors. We will first use the “str” function to identify factor variables and then remove them from the dataset. We also need to remove in examples that are missing data so we use the “na.omit” function for this. Below is the code
survey$Clap<-NULL
survey$W.Hnd<-NULL
survey$Fold<-NULL
survey$Exer<-NULL
survey$Smoke<-NULL
survey$M.I<-NULL
survey<-na.omit(survey)We now need to check for collinearity using the “corrplot.mixed” function form the “corrplot” package.
pc<-cor(survey[,2:5])
corrplot.mixed(pc)
corrplot.mixed(pc)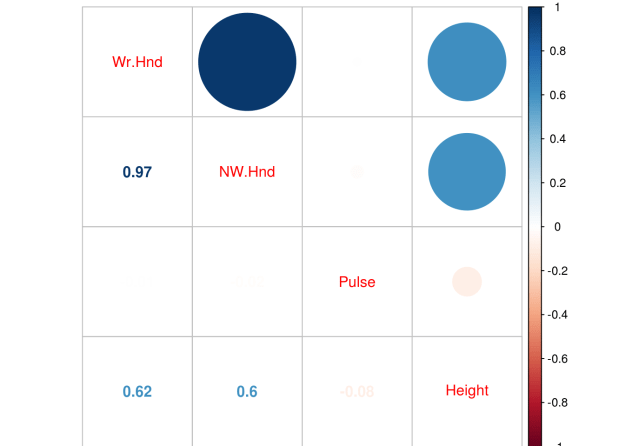
We have an extreme correlation between “We.Hnd” and “NW.Hnd” this makes sense because people’s hands are normally the same size. Since this blog post is a demonstration of logistic regression we will not worry about this too much.
We now need to divide our dataset into a train and a test set. We set the seed for. First, we need to make a variable that we call “ind” that is randomly assigned 70% of the number of rows of survey 1 and 30% 2. We then subset the “train” dataset by taking all rows that are 1’s based on the “ind” variable and we create the “test” dataset for all the rows that line up with 2 in the “ind” variable. This means our data split is 70% train and 30% test. Below is the code
set.seed(123)
ind<-sample(2,nrow(survey),replace=T,prob = c(0.7,0.3))
train<-survey[ind==1,]
test<-survey[ind==2,]We now make our model. We use the “glm” function for logistic regression. We set the family argument to “binomial”. Next, we look at the results as well as the odds ratios.
fit<-glm(Sex~.,family=binomial,train)
summary(fit)##
## Call:
## glm(formula = Sex ~ ., family = binomial, data = train)
##
## Deviance Residuals:
## Min 1Q Median 3Q Max
## -1.9875 -0.5466 -0.1395 0.3834 3.4443
##
## Coefficients:
## Estimate Std. Error z value Pr(>|z|)
## (Intercept) -46.42175 8.74961 -5.306 1.12e-07 ***
## Wr.Hnd -0.43499 0.66357 -0.656 0.512
## NW.Hnd 1.05633 0.70034 1.508 0.131
## Pulse -0.02406 0.02356 -1.021 0.307
## Height 0.21062 0.05208 4.044 5.26e-05 ***
## Age 0.00894 0.05368 0.167 0.868
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
##
## (Dispersion parameter for binomial family taken to be 1)
##
## Null deviance: 169.14 on 122 degrees of freedom
## Residual deviance: 81.15 on 117 degrees of freedom
## AIC: 93.15
##
## Number of Fisher Scoring iterations: 6exp(coef(fit))## (Intercept) Wr.Hnd NW.Hnd Pulse Height
## 6.907034e-21 6.472741e-01 2.875803e+00 9.762315e-01 1.234447e+00
## Age
## 1.008980e+00The results indicate that only height is useful in predicting if someone is a male or female. The second piece of code shares the odds ratios. The odds ratio tell how a one unit increase in the independent variable leads to an increase in the odds of being male in our model. For example, for every one unit increase in height there is a 1.23 increase in the odds of a particular example being male.
We now need to see how well our model does on the train and test dataset. We first capture the probabilities and save them to the train dataset as “probs”. Next we create a “predict” variable and place the string “Female” in the same number of rows as are in the “train” dataset. Then we rewrite the “predict” variable by changing any example that has a probability above 0.5 as “Male”. Then we make a table of our results to see the number correct, false positives/negatives. Lastly, we calculate the accuracy rate. Below is the code.
train$probs<-predict(fit, type = 'response')
train$predict<-rep('Female',123)
train$predict[train$probs>0.5]<-"Male"
table(train$predict,train$Sex)##
## Female Male
## Female 61 7
## Male 7 48mean(train$predict==train$Sex)## [1] 0.8861789Despite the weaknesses of the model with so many insignificant variables it is surprisingly accurate at 88.6%. Let’s see how well we do on the “test” dataset.
test$prob<-predict(fit,newdata = test, type = 'response')
test$predict<-rep('Female',46)
test$predict[test$prob>0.5]<-"Male"
table(test$predict,test$Sex)##
## Female Male
## Female 17 3
## Male 0 26mean(test$predict==test$Sex)## [1] 0.9347826As you can see, we do even better on the test set with an accuracy of 93.4%. Our model is looking pretty good and height is an excellent predictor of sex which makes complete sense. However, in the next post we will use cross-validation and the ROC plot to further assess the quality of it.

Probability,Odds, and Odds Ratio
In logistic regression, there are three terms that are used frequently but can be confusing if they are not thoroughly explained. These three terms are probability, odds, and odds ratio. In this post, we will look at these three terms and provide an explanation of them.
Probability
Probability is probably (no pun intended) the easiest of these three terms to understand. Probability is simply the likelihood that a certain event will happen. To calculate the probability in the traditional sense you need to know the number of events and outcomes to find the probability.
Bayesian probability uses prior probabilities to develop a posterior probability based on new evidence. For example, at one point during Super Bowl LI the Atlanta Falcons had a 99.7% chance of winning. This was base don such factors as the number points they were ahead and the time remaining. As these changed, so did the probability of them winning. yet the Patriots still found a way to win with less than a 1% chance
Bayesian probability was also used for predicting who would win the 2016 US presidential race. It is important to remember that probability is an expression of confidence and not a guarantee as we saw in both examples.
Odds
Odds are the expression of relative probabilities. Odds are calculated using the following equation
probability of the event ⁄ 1 – probability of the event
For example, at one point during Super Bowl LI the odds of the Atlanta Falcons winning were as follows
0.997 ⁄ 1 – 0.997 = 332
This can be interpreted as the odds being 332 to 1! This means that Atlanta was 332 times more likely to win the Super Bowl then loss the Super Bowl.
Odds are commonly used in gambling and this is probably (again no pun intended) where most of us have heard the term before. The odds is just an extension of probabilities and they are most commonly expressed as a fraction such as one in four, etc.
Odds Ratio
A ratio is the comparison of two numbers and indicates how many times one number is contained or contains another number. For example, a ration of boys to girls is 5 to 1 it means that there are five boys for every one girl.
By extension odds ratio is the comparison of two different odds. For example, if the odds of Team A making the playoffs is 45% and the odds of Team B making the playoffs is 35% the odds ratio is calculated as follows.
0.45 ⁄ 0.35 = 1.28
Team A is 1.28 more likely to make the playoffs then Team B.
The value of the odds and the odds ratio can sometimes be the same. Below is the odds ratio of the Atlanta Falcons winning and the New Patriots winning Super Bowl LI
0.997⁄ 0.003 = 332
As such there is little difference between odds and odds ratio except that odds ratio is the ratio of two odds ratio. As you can tell, there is a lot of confusion about this for the average person. However, understanding these terms is critical to the application of logistic regression.
Best Subset Regression in R
In this post, we will take a look at best subset regression. Best subset regression fits a model for all possible feature or variable combinations and the decision for the most appropriate model is made by the analyst based on judgment or some statistical criteria.
Best subset regression is an alternative to both Forward and Backward stepwise regression. Forward stepwise selection adds one variable at a time based on the lowest residual sum of squares until no more variables continue to lower the residual sum of squares. Backward stepwise regression starts with all variables in the model and removes variables one at a time. The concern with stepwise methods is they can produce biased regression coefficients, conflicting models, and inaccurate confidence intervals.
Best subset regression bypasses these weaknesses of stepwise models by creating all models possible and then allowing you to assess which variables should be included in your final model. The one drawback to best subset is that a large number of variables means a large number of potential models, which can make it difficult to make a decision among several choices.
In this post, we will use the “Fair” dataset from the “Ecdat” package to predict marital satisfaction based on age, Sex, the presence of children, years married, religiosity, education, occupation, and the number of affairs in the past year. Below is some initial code.
library(leaps);library(Ecdat);library(car);library(lmtest)data(Fair)
We begin our analysis by building the initial model with all variables in it. Below is the code
fit<-lm(rate~.,Fair)
summary(fit)##
## Call:
## lm(formula = rate ~ ., data = Fair)
##
## Residuals:
## Min 1Q Median 3Q Max
## -3.2049 -0.6661 0.2298 0.7705 2.2292
##
## Coefficients:
## Estimate Std. Error t value Pr(>|t|)
## (Intercept) 3.522875 0.358793 9.819 < 2e-16 ***
## sexmale -0.062281 0.099952 -0.623 0.53346
## age -0.009683 0.007548 -1.283 0.20005
## ym -0.019978 0.013887 -1.439 0.15079
## childyes -0.206976 0.116227 -1.781 0.07546 .
## religious 0.042142 0.037705 1.118 0.26416
## education 0.068874 0.021153 3.256 0.00119 **
## occupation -0.015606 0.029602 -0.527 0.59825
## nbaffairs -0.078812 0.013286 -5.932 5.09e-09 ***
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
##
## Residual standard error: 1.03 on 592 degrees of freedom
## Multiple R-squared: 0.1405, Adjusted R-squared: 0.1289
## F-statistic: 12.1 on 8 and 592 DF, p-value: 4.487e-16The initial results are already interesting even though the r-square is low. When couples have children the have less marital satisfaction than couples without children when controlling for the other factors and this is the strongest regression weight. In addition, the more education a person has there is an increase in marital satisfaction. Lastly, as the number of affairs increases there is also a decrease in marital satisfaction. Keep in mind that the “rate” variable goes from 1 to 5 with one meaning a terrible marriage to five being a great one. The mean marital satisfaction was 3.52 when controlling for the other variables.
We will now create our subset models. Below is the code.
sub.fit<-regsubsets(rate~.,Fair)
best.summary<-summary(sub.fit)In the code above we create the sub models using the “regsubsets” function from the “leaps” package and saved it in the variable called “sub.fit”. We then saved the summary of “sub.fit” in the variable “best.summary”. We will use the “best.summary” “sub.fit variables several times to determine which model to use.
There are many different ways to assess the model. We will use the following statistical methods that come with the results from the “regsubset” function.
- Mallow’ Cp
- Bayesian Information Criteria
We will make two charts for each of the criteria above. The plot to the left will explain how many features to include in the model. The plot to the right will tell you which variables to include. It is important to note that for both of these methods, the lower the score the better the model. Below is the code for Mallow’s Cp.
par(mfrow=c(1,2))
plot(best.summary$cp)
plot(sub.fit,scale = "Cp")
The plot on the left suggests that a four feature model is the most appropriate. However, this chart does not tell me which four features. The chart on the right is read in reverse order. The high numbers are at the bottom and the low numbers are at the top when looking at the y-axis. Knowing this, we can conclude that the most appropriate variables to include in the model are age, children presence, education, and number of affairs. Below are the results using the Bayesian Information Criterion
par(mfrow=c(1,2))
plot(best.summary$bic)
plot(sub.fit,scale = "bic")
These results indicate that a three feature model is appropriate. The variables or features are years married, education, and number of affairs. Presence of children was not considered beneficial. Since our original model and Mallow’s Cp indicated that presence of children was significant we will include it for now.
Below is the code for the model based on the subset regression.
fit2<-lm(rate~age+child+education+nbaffairs,Fair)
summary(fit2)##
## Call:
## lm(formula = rate ~ age + child + education + nbaffairs, data = Fair)
##
## Residuals:
## Min 1Q Median 3Q Max
## -3.2172 -0.7256 0.1675 0.7856 2.2713
##
## Coefficients:
## Estimate Std. Error t value Pr(>|t|)
## (Intercept) 3.861154 0.307280 12.566 < 2e-16 ***
## age -0.017440 0.005057 -3.449 0.000603 ***
## childyes -0.261398 0.103155 -2.534 0.011531 *
## education 0.058637 0.017697 3.313 0.000978 ***
## nbaffairs -0.084973 0.012830 -6.623 7.87e-11 ***
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
##
## Residual standard error: 1.029 on 596 degrees of freedom
## Multiple R-squared: 0.1352, Adjusted R-squared: 0.1294
## F-statistic: 23.29 on 4 and 596 DF, p-value: < 2.2e-16The results look ok. The older a person is the less satisfied they are with their marriage. If children are present the marriage is less satisfying. The more educated the more satisfied they are. Lastly, the higher the number of affairs indicate less marital satisfaction. However, before we get excited we need to check for collinearity and homoscedasticity. Below is the code
vif(fit2)## age child education nbaffairs
## 1.249430 1.228733 1.023722 1.014338No issues with collinearity.For vif values above 5 or 10 indicate a problem. Let’s check for homoscedasticity
par(mfrow=c(2,2))
plot(fit2)
The normal qqplot and residuals vs leverage plot can be used for locating outliers. The residual vs fitted and the scale-location plot do not look good as there appears to be a pattern in the dispersion which indicates homoscedasticity. To confirm this we will use Breusch-Pagan test from the “lmtest” package. Below is the code
bptest(fit2)##
## studentized Breusch-Pagan test
##
## data: fit2
## BP = 16.238, df = 4, p-value = 0.002716There you have it. Our model violates the assumption of homoscedasticity. However, this model was developed for demonstration purpose to provide an example of subset regression.
Linear Discriminant Analysis in R
In this post we will look at an example of linear discriminant analysis (LDA). LDA is used to develop a statistical model that classifies examples in a dataset. In the example in this post, we will use the “Star” dataset from the “Ecdat” package. What we will do is try to predict the type of class the students learned in (regular, small, regular with aide) using their math scores, reading scores, and the teaching experience of the teacher. Below is the initial code
library(Ecdat)library(MASS)data(Star)
We first need to examine the data by using the “str” function
str(Star)## 'data.frame': 5748 obs. of 8 variables:
## $ tmathssk: int 473 536 463 559 489 454 423 500 439 528 ...
## $ treadssk: int 447 450 439 448 447 431 395 451 478 455 ...
## $ classk : Factor w/ 3 levels "regular","small.class",..: 2 2 3 1 2 1 3 1 2 2 ...
## $ totexpk : int 7 21 0 16 5 8 17 3 11 10 ...
## $ sex : Factor w/ 2 levels "girl","boy": 1 1 2 2 2 2 1 1 1 1 ...
## $ freelunk: Factor w/ 2 levels "no","yes": 1 1 2 1 2 2 2 1 1 1 ...
## $ race : Factor w/ 3 levels "white","black",..: 1 2 2 1 1 1 2 1 2 1 ...
## $ schidkn : int 63 20 19 69 79 5 16 56 11 66 ...
## - attr(*, "na.action")=Class 'omit' Named int [1:5850] 1 4 6 7 8 9 10 15 16 17 ...
## .. ..- attr(*, "names")= chr [1:5850] "1" "4" "6" "7" ...We will use the following variables
- dependent variable = classk (class type)
- independent variable = tmathssk (Math score)
- independent variable = treadssk (Reading score)
- independent variable = totexpk (Teaching experience)
We now need to examine the data visually by looking at histograms for our independent variables and a table for our dependent variable
hist(Star$tmathssk)
hist(Star$treadssk)
hist(Star$totexpk)
prop.table(table(Star$classk))##
## regular small.class regular.with.aide
## 0.3479471 0.3014962 0.3505567The data mostly looks good. The results of the “prop.table” function will help us when we develop are training and testing datasets. The only problem is with the “totexpk” variable. IT is not anywhere near to be normally distributed. TO deal with this we will use the square root for teaching experience. Below is the code
star.sqrt<-Star
star.sqrt$totexpk.sqrt<-sqrt(star.sqrt$totexpk)
hist(sqrt(star.sqrt$totexpk))
Much better. We now need to check the correlation among the variables as well and we will use the code below.
cor.star<-data.frame(star.sqrt$tmathssk,star.sqrt$treadssk,star.sqrt$totexpk.sqrt)
cor(cor.star)## star.sqrt.tmathssk star.sqrt.treadssk
## star.sqrt.tmathssk 1.00000000 0.7135489
## star.sqrt.treadssk 0.71354889 1.0000000
## star.sqrt.totexpk.sqrt 0.08647957 0.1045353
## star.sqrt.totexpk.sqrt
## star.sqrt.tmathssk 0.08647957
## star.sqrt.treadssk 0.10453533
## star.sqrt.totexpk.sqrt 1.00000000None of the correlations are too bad. We can now develop our model using linear discriminant analysis. First, we need to scale are scores because the test scores and the teaching experience are measured differently. Then, we need to divide our data into a train and test set as this will allow us to determine the accuracy of the model. Below is the code.
star.sqrt$tmathssk<-scale(star.sqrt$tmathssk)
star.sqrt$treadssk<-scale(star.sqrt$treadssk)
star.sqrt$totexpk.sqrt<-scale(star.sqrt$totexpk.sqrt)
train.star<-star.sqrt[1:4000,]
test.star<-star.sqrt[4001:5748,]Now we develop our model. In the code before the “prior” argument indicates what we expect the probabilities to be. In our data the distribution of the the three class types is about the same which means that the apriori probability is 1/3 for each class type.
train.lda<-lda(classk~tmathssk+treadssk+totexpk.sqrt, data =
train.star,prior=c(1,1,1)/3)
train.lda## Call:
## lda(classk ~ tmathssk + treadssk + totexpk.sqrt, data = train.star,
## prior = c(1, 1, 1)/3)
##
## Prior probabilities of groups:
## regular small.class regular.with.aide
## 0.3333333 0.3333333 0.3333333
##
## Group means:
## tmathssk treadssk totexpk.sqrt
## regular -0.04237438 -0.05258944 -0.05082862
## small.class 0.13465218 0.11021666 -0.02100859
## regular.with.aide -0.05129083 -0.01665593 0.09068835
##
## Coefficients of linear discriminants:
## LD1 LD2
## tmathssk 0.89656393 -0.04972956
## treadssk 0.04337953 0.56721196
## totexpk.sqrt -0.49061950 0.80051026
##
## Proportion of trace:
## LD1 LD2
## 0.7261 0.2739The printout is mostly readable. At the top is the actual code used to develop the model followed by the probabilities of each group. The next section shares the means of the groups. The coefficients of linear discriminants are the values used to classify each example. The coefficients are similar to regression coefficients. The computer places each example in both equations and probabilities are calculated. Whichever class has the highest probability is the winner. In addition, the higher the coefficient the more weight it has. For example, “tmathssk” is the most influential on LD1 with a coefficient of 0.89.
The proportion of trace is similar to principal component analysis
Now we will take the trained model and see how it does with the test set. We create a new model called “predict.lda” and use are “train.lda” model and the test data called “test.star”
predict.lda<-predict(train.lda,newdata = test.star)We can use the “table” function to see how well are model has done. We can do this because we actually know what class our data is beforehand because we divided the dataset. What we need to do is compare this to what our model predicted. Therefore, we compare the “classk” variable of our “test.star” dataset with the “class” predicted by the “predict.lda” model.
table(test.star$classk,predict.lda$class)##
## regular small.class regular.with.aide
## regular 155 182 249
## small.class 145 198 174
## regular.with.aide 172 204 269The results are pretty bad. For example, in the first row called “regular” we have 155 examples that were classified as “regular” and predicted as “regular” by the model. In rhe next column, 182 examples that were classified as “regular” but predicted as “small.class”, etc. To find out how well are model did you add together the examples across the diagonal from left to right and divide by the total number of examples. Below is the code
(155+198+269)/1748## [1] 0.3558352Only 36% accurate, terrible but ok for a demonstration of linear discriminant analysis. Since we only have two-functions or two-dimensions we can plot our model. Below I provide a visual of the first 50 examples classified by the predict.lda model.
plot(predict.lda$x[1:50])
text(predict.lda$x[1:50],as.character(predict.lda$class[1:50]),col=as.numeric(predict.lda$class[1:100]))
abline(h=0,col="blue")
abline(v=0,col="blue")
The first function, which is the vertical line, doesn’t seem to discriminant anything as it off to the side and not separating any of the data. However, the second function, which is the horizontal one, does a good of dividing the “regular.with.aide” from the “small.class”. Yet, there are problems with distinguishing the class “regular” from either of the other two groups. In order improve our model we need additional independent variables to help to distinguish the groups in the dependent variable.
Generalized Additive Models in R

In our example, we will use the “Auto” dataset from the “ISLR” package and use the variables “mpg”,“displacement”,“horsepower”, and “weight” to predict “acceleration”. We will also use the “mgcv” package. Below is some initial code to begin the analysis
library(mgcv)library(ISLR) data(Auto)
We will now make the model we want to understand the response of “acceleration” to the explanatory variables of “mpg”,“displacement”,“horsepower”, and “weight”. After setting the model we will examine the summary. Below is the code
model1<-gam(acceleration~s(mpg)+s(displacement)+s(horsepower)+s(weight),data=Auto)
summary(model1)##
## Family: gaussian
## Link function: identity
##
## Formula:
## acceleration ~ s(mpg) + s(displacement) + s(horsepower) + s(weight)
##
## Parametric coefficients:
## Estimate Std. Error t value Pr(>|t|)
## (Intercept) 15.54133 0.07205 215.7 <2e-16 ***
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
##
## Approximate significance of smooth terms:
## edf Ref.df F p-value
## s(mpg) 6.382 7.515 3.479 0.00101 **
## s(displacement) 1.000 1.000 36.055 4.35e-09 ***
## s(horsepower) 4.883 6.006 70.187 < 2e-16 ***
## s(weight) 3.785 4.800 41.135 < 2e-16 ***
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
##
## R-sq.(adj) = 0.733 Deviance explained = 74.4%
## GCV = 2.1276 Scale est. = 2.0351 n = 392All of the explanatory variables are significant and the adjust r-squared is .73 which is excellent. edf stands for “effective degrees of freedom”. This modified version of the degree of freedoms is due to the smoothing process in the model. GCV stands for generalized cross-validation and this number is useful when comparing models. The model with the lowest number is the better model.
We can also examine the model visually by using the “plot” function. This will allow us to examine if the curvature fitted by the smoothing process was useful or not for each variable. Below is the code.
plot(model1)



We can also look at a 3d graph that includes the linear predictor as well as the two strongest predictors. This is done with the “vis.gam” function. Below is the code
vis.gam(model1)
If multiple models are developed. You can compare the GCV values to determine which model is the best. In addition, another way to compare models is with the “AIC” function. In the code below, we will create an additional model that includes “year” compare the GCV scores and calculate the AIC. Below is the code.
model2<-gam(acceleration~s(mpg)+s(displacement)+s(horsepower)+s(weight)+s(year),data=Auto)
summary(model2)##
## Family: gaussian
## Link function: identity
##
## Formula:
## acceleration ~ s(mpg) + s(displacement) + s(horsepower) + s(weight) +
## s(year)
##
## Parametric coefficients:
## Estimate Std. Error t value Pr(>|t|)
## (Intercept) 15.54133 0.07203 215.8 <2e-16 ***
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
##
## Approximate significance of smooth terms:
## edf Ref.df F p-value
## s(mpg) 5.578 6.726 2.749 0.0106 *
## s(displacement) 2.251 2.870 13.757 3.5e-08 ***
## s(horsepower) 4.936 6.054 66.476 < 2e-16 ***
## s(weight) 3.444 4.397 34.441 < 2e-16 ***
## s(year) 1.682 2.096 0.543 0.6064
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
##
## R-sq.(adj) = 0.733 Deviance explained = 74.5%
## GCV = 2.1368 Scale est. = 2.0338 n = 392#model1 GCV
model1$gcv.ubre## GCV.Cp
## 2.127589#model2 GCV
model2$gcv.ubre## GCV.Cp
## 2.136797As you can see, the second model has a higher GCV score when compared to the first model. This indicates that the first model is a better choice. This makes sense because in the second model the variable “year” is not significant. To confirm this we will calculate the AIC scores using the AIC function.
AIC(model1,model2)## df AIC
## model1 18.04952 1409.640
## model2 19.89068 1411.156Again, you can see that model1 s better due to its fewer degrees of freedom and slightly lower AIC score.
Conclusion
Using GAMs is most common for exploring potential relationships in your data. This is stated because they are difficult to interpret and to try and summarize. Therefore, it is normally better to develop a generalized linear model over a GAM due to the difficulty in understanding what the data is trying to tell you when using GAMs.























